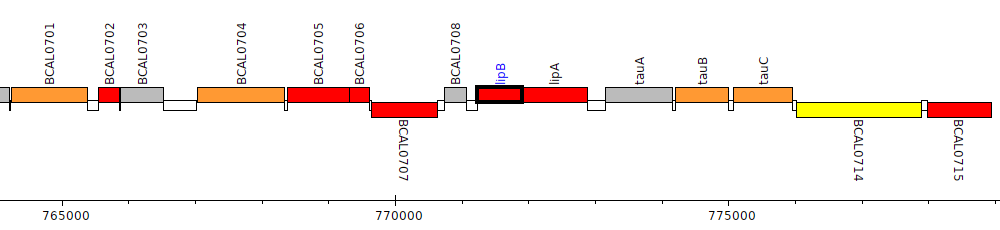
Burkholderia cenocepacia J2315, BCAL0709 (lipB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009249 | protein lipoylation |
Inferred from Sequence Model
Term mapped from: InterPro:cd16444
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006464 | cellular protein modification process |
Inferred from Sequence Model
Term mapped from: InterPro:PS51733
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0033819 | lipoyl(octanoyl) transferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd16444
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | lipoate biosynthesis and incorporation (yeast) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj00785 | Lipoic acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | lipoate biosynthesis and incorporation III (Bacillus) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00785 | Lipoic acid metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| ProSitePatterns | PS01313 | Lipoate-protein ligase B signature. | IPR020605 | Octanoyltransferase, conserved site | 65 | 80 | - |
| ProSiteProfiles | PS51733 | Biotinyl protein ligase (BPL) and lipoyl protein ligase (LPL) catalytic domain profile. | IPR004143 | Biotinyl protein ligase (BPL) and lipoyl protein ligase (LPL), catalytic domain | 25 | 206 | 57.338 |
| CDD | cd16444 | LipB | IPR000544 | Octanoyltransferase | 2 | 202 | 4.59764E-87 |
| PIRSF | PIRSF016262 | IPR000544 | Octanoyltransferase | 1 | 217 | 1.1E-57 | |
| Hamap | MF_00013 | Octanoyltransferase [lipB]. | IPR000544 | Octanoyltransferase | 1 | 205 | 35.047 |
| TIGRFAM | TIGR00214 | lipB: lipoyl(octanoyl) transferase | IPR000544 | Octanoyltransferase | 19 | 200 | 2.5E-58 |
| Gene3D | G3DSA:3.30.930.10 | 1 | 203 | 1.8E-58 | |||
| SUPERFAMILY | SSF55681 | 3 | 203 | 7.21E-60 | |||
| Pfam | PF03099 | Biotin/lipoate A/B protein ligase family | IPR004143 | Biotinyl protein ligase (BPL) and lipoyl protein ligase (LPL), catalytic domain | 56 | 158 | 2.1E-6 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.