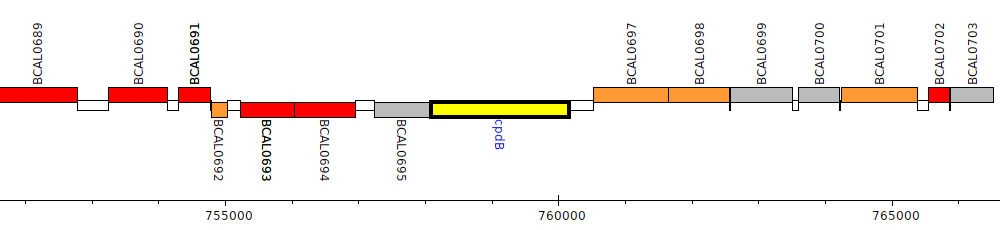
Burkholderia cenocepacia J2315, BCAL0696 (cpdB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016787 | hydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00149
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0046872 | metal ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS00786
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009166 | nucleotide catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF02872
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016788 | hydrolase activity, acting on ester bonds |
Inferred from Sequence Model
Term mapped from: InterPro:PS00786
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000166 | nucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS00786
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj00230 | Purine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00240 | Pyrimidine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00149 | Calcineurin-like phosphoesterase | IPR004843 | Calcineurin-like phosphoesterase domain, ApaH type | 56 | 311 | 1.3E-4 |
| PRINTS | PR01607 | Apyrase family signature | IPR006179 | 5'-Nucleotidase/apyrase | 572 | 591 | 2.8E-28 |
| Gene3D | G3DSA:3.90.780.10 | IPR036907 | 5'-Nucleotidase, C-terminal domain superfamily | 425 | 631 | 1.5E-53 | |
| ProSiteProfiles | PS51318 | Twin arginine translocation (Tat) signal profile. | IPR006311 | Twin-arginine translocation pathway, signal sequence | 1 | 44 | 7.891 |
| SUPERFAMILY | SSF56300 | 48 | 407 | 1.19E-75 | |||
| PRINTS | PR01607 | Apyrase family signature | IPR006179 | 5'-Nucleotidase/apyrase | 482 | 505 | 2.8E-28 |
| PRINTS | PR01607 | Apyrase family signature | IPR006179 | 5'-Nucleotidase/apyrase | 266 | 283 | 2.8E-28 |
| Gene3D | G3DSA:3.60.21.10 | IPR029052 | Metallo-dependent phosphatase-like | 47 | 409 | 3.3E-108 | |
| ProSitePatterns | PS00786 | 5'-nucleotidase signature 2. | IPR006146 | 5'-Nucleotidase, conserved site | 138 | 149 | - |
| PRINTS | PR01607 | Apyrase family signature | IPR006179 | 5'-Nucleotidase/apyrase | 294 | 317 | 2.8E-28 |
| PRINTS | PR01607 | Apyrase family signature | IPR006179 | 5'-Nucleotidase/apyrase | 338 | 358 | 2.8E-28 |
| SUPERFAMILY | SSF55816 | IPR036907 | 5'-Nucleotidase, C-terminal domain superfamily | 408 | 631 | 5.49E-39 | |
| Pfam | PF02872 | 5'-nucleotidase, C-terminal domain | IPR008334 | 5'-Nucleotidase, C-terminal | 425 | 597 | 5.4E-22 |
| PRINTS | PR01607 | Apyrase family signature | IPR006179 | 5'-Nucleotidase/apyrase | 50 | 68 | 2.8E-28 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.