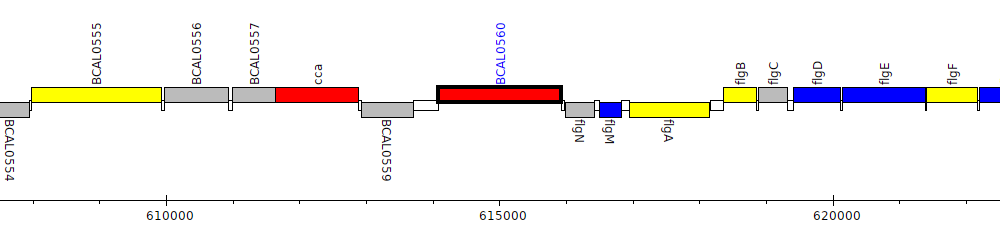
Burkholderia cenocepacia J2315, BCAL0560
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016787 | hydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01321
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| ProSitePatterns | PS00491 | Aminopeptidase P and proline dipeptidase signature. | IPR001131 | Peptidase M24B, X-Pro dipeptidase/aminopeptidase P, conserved site | 505 | 517 | - |
| Gene3D | G3DSA:3.40.350.10 | IPR029149 | Creatinase/Aminopeptidase P/Spt16, N-terminal | 181 | 322 | 4.8E-6 | |
| Pfam | PF16189 | Creatinase/Prolidase N-terminal domain | 183 | 340 | 6.8E-12 | ||
| SUPERFAMILY | SSF55920 | IPR036005 | Creatinase/aminopeptidase-like | 339 | 592 | 5.63E-48 | |
| Gene3D | G3DSA:3.40.350.10 | IPR029149 | Creatinase/Aminopeptidase P/Spt16, N-terminal | 18 | 174 | 1.4E-25 | |
| Pfam | PF00557 | Metallopeptidase family M24 | IPR000994 | Peptidase M24 | 345 | 559 | 4.9E-43 |
| Gene3D | G3DSA:3.90.230.10 | 336 | 612 | 2.6E-65 | |||
| Pfam | PF01321 | Creatinase/Prolidase N-terminal domain | IPR000587 | Creatinase, N-terminal | 26 | 158 | 4.2E-9 |
| SUPERFAMILY | SSF53092 | IPR029149 | Creatinase/Aminopeptidase P/Spt16, N-terminal | 24 | 159 | 5.75E-6 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.