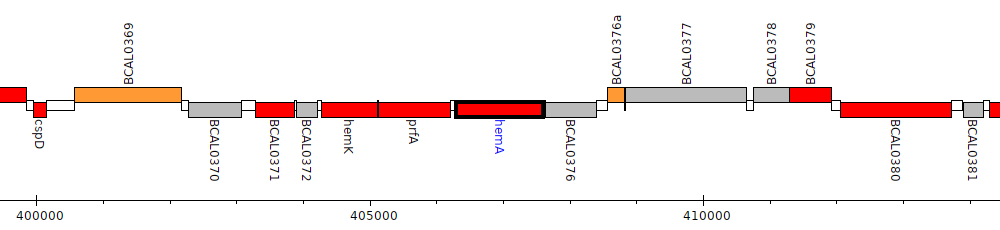
Burkholderia cenocepacia J2315, BCAL0375 (hemA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00087
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008883 | glutamyl-tRNA reductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00087
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0033014 | tetrapyrrole biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00087
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0050661 | NADP binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00087
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | tetrapyrrole biosynthesis I (from glutamate) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00860 | Porphyrin and chlorophyll metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00860 | Porphyrin and chlorophyll metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.30.460.30 | IPR036343 | Glutamyl-tRNA reductase, N-terminal domain superfamily | 1 | 163 | 8.4E-54 | |
| SUPERFAMILY | SSF69075 | IPR036453 | Glutamyl tRNA-reductase dimerization domain superfamily | 322 | 422 | 4.45E-28 | |
| Gene3D | G3DSA:3.40.50.720 | 166 | 317 | 1.5E-44 | |||
| SUPERFAMILY | SSF51735 | IPR036291 | NAD(P)-binding domain superfamily | 168 | 320 | 3.04E-36 | |
| TIGRFAM | TIGR01035 | hemA: glutamyl-tRNA reductase | IPR000343 | Tetrapyrrole biosynthesis, glutamyl-tRNA reductase | 4 | 413 | 6.1E-126 |
| Pfam | PF00745 | Glutamyl-tRNAGlu reductase, dimerisation domain | IPR015896 | Tetrapyrrole biosynthesis, glutamyl-tRNA reductase, dimerisation domain | 325 | 420 | 1.1E-26 |
| CDD | cd05213 | NAD_bind_Glutamyl_tRNA_reduct | 3 | 323 | 1.10595E-124 | ||
| ProSitePatterns | PS00747 | Glutamyl-tRNA reductase signature. | IPR018214 | Glutamyl-tRNA reductase, conserved site | 104 | 127 | - |
| PIRSF | PIRSF000445 | IPR000343 | Tetrapyrrole biosynthesis, glutamyl-tRNA reductase | 1 | 432 | 4.8E-153 | |
| SUPERFAMILY | SSF69742 | IPR036343 | Glutamyl-tRNA reductase, N-terminal domain superfamily | 1 | 163 | 8.5E-46 | |
| Pfam | PF01488 | Shikimate / quinate 5-dehydrogenase | IPR006151 | Quinate/shikimate 5-dehydrogenase/glutamyl-tRNA reductase | 177 | 310 | 2.8E-45 |
| Hamap | MF_00087 | Glutamyl-tRNA reductase [hemA]. | IPR000343 | Tetrapyrrole biosynthesis, glutamyl-tRNA reductase | 2 | 419 | 29.483 |
| Pfam | PF05201 | Glutamyl-tRNAGlu reductase, N-terminal domain | IPR015895 | Tetrapyrrole biosynthesis, glutamyl-tRNA reductase, N-terminal | 6 | 161 | 5.3E-48 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.