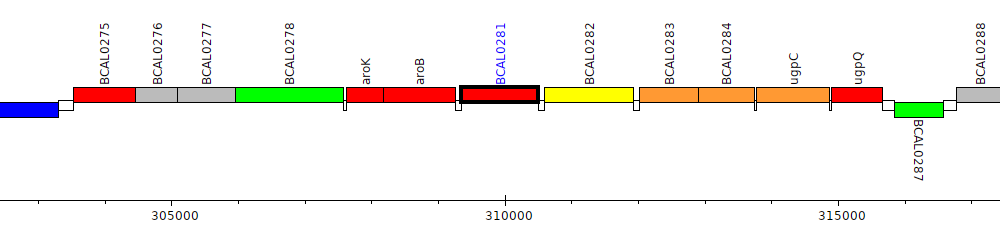
Burkholderia cenocepacia J2315, BCAL0281
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016793 | triphosphoric monoester hydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01353
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj00230 | Purine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd00077 | HDc | IPR003607 | HD/PDEase domain | 72 | 215 | 2.09761E-9 |
| Pfam | PF01966 | HD domain | IPR006674 | HD domain | 74 | 204 | 1.9E-15 |
| Hamap | MF_01212 | Deoxyguanosinetriphosphate triphosphohydrolase-like protein. | IPR023023 | dNTP triphosphohydrolase, type 2 | 14 | 378 | 38.355 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 18 | 43 | - | ||
| SMART | SM00471 | IPR003607 | HD/PDEase domain | 70 | 214 | 6.0E-11 | |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 24 | 43 | - | ||
| TIGRFAM | TIGR01353 | dGTP_triPase: putative dGTPase | IPR006261 | dNTP triphosphohydrolase | 37 | 170 | 1.3E-58 |
| Pfam | PF13286 | Phosphohydrolase-associated domain | IPR026875 | Phosphohydrolase-associated domain | 291 | 376 | 1.0E-27 |
| ProSiteProfiles | PS51831 | HD domain profile. | IPR003607 | HD/PDEase domain | 74 | 205 | 23.385 |
| Gene3D | G3DSA:1.10.3210.10 | 5 | 379 | 3.4E-135 | |||
| SUPERFAMILY | SSF109604 | 25 | 379 | 2.51E-92 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.