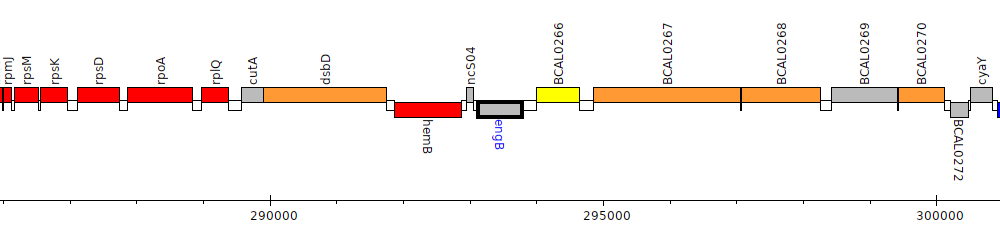
Burkholderia cenocepacia J2315, BCAL0265 (engB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0005525 | GTP binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd01876
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd01876 | YihA_EngB | IPR030393 | EngB-type guanine nucleotide-binding (G) domain | 27 | 205 | 8.15084E-79 |
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 14 | 205 | 7.53E-32 | |
| ProSiteProfiles | PS51706 | EngB-type guanine nucleotide-binding (G) domain profile. | IPR030393 | EngB-type guanine nucleotide-binding (G) domain | 24 | 207 | 36.424 |
| Hamap | MF_00321 | Probable GTP-binding protein EngB [engB]. | IPR019987 | GTP-binding protein, ribosome biogenesis, YsxC | 13 | 205 | 25.734 |
| Pfam | PF01926 | 50S ribosome-binding GTPase | IPR006073 | GTP binding domain | 27 | 149 | 5.1E-19 |
| TIGRFAM | TIGR03598 | GTPase_YsxC: ribosome biogenesis GTP-binding protein YsxC | IPR019987 | GTP-binding protein, ribosome biogenesis, YsxC | 8 | 201 | 4.8E-64 |
| Gene3D | G3DSA:3.40.50.300 | 1 | 219 | 4.5E-62 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.