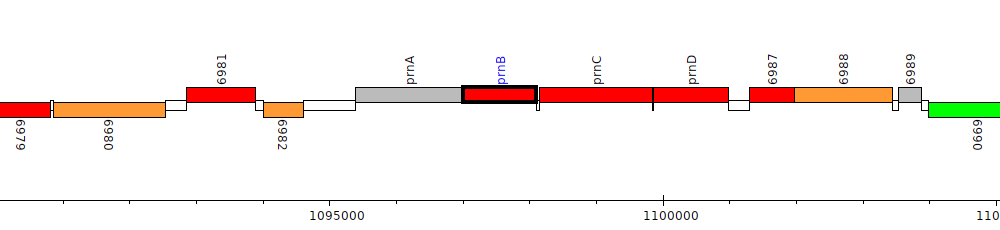
Burkholderia cenocepacia MC0-3, Bcenmc03_6984 (prnB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0019441 | tryptophan catabolic process to kynurenine |
Inferred from Sequence Model
Term mapped from: InterPro:SSF140959
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0046872 | metal ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF140959
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0020037 | heme binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF140959
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00380 | Tryptophan metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:1.20.58.480 | 134 | 361 | 4.8E-82 | |||
| Pfam | PF08933 | Domain of unknown function (DUF1864) | IPR015029 | Protein of unknown function DUF1864 | 14 | 353 | 3.3E-110 |
| Gene3D | G3DSA:1.20.58.1320 | 1 | 118 | 1.8E-46 | |||
| SUPERFAMILY | SSF140959 | IPR037217 | Tryptophan/Indoleamine 2,3-dioxygenase-like | 20 | 351 | 8.24E-57 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.