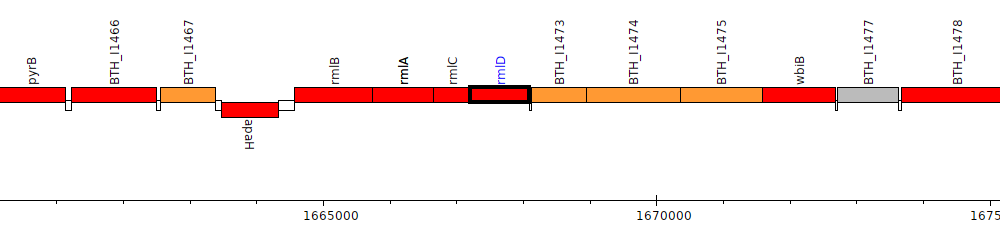
Burkholderia thailandensis E264 ATCC 700388, BTH_I1472 (rmlD)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bte01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte00523 | Polyketide sugar unit biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte00521 | Streptomycin biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF51735 | IPR036291 | NAD(P)-binding domain superfamily | 1 | 295 | 3.19E-94 | |
| Gene3D | G3DSA:3.90.25.10 | 158 | 287 | 3.2E-104 | |||
| Gene3D | G3DSA:3.40.50.720 | 3 | 282 | 3.2E-104 | |||
| Pfam | PF04321 | RmlD substrate binding domain | IPR029903 | RmlD-like substrate binding domain | 1 | 295 | 4.3E-110 |
| TIGRFAM | TIGR01214 | rmlD: dTDP-4-dehydrorhamnose reductase | IPR005913 | dTDP-4-dehydrorhamnose reductase family | 2 | 292 | 2.0E-99 |
| CDD | cd05254 | dTDP_HR_like_SDR_e | 2 | 285 | 1.07918E-85 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.