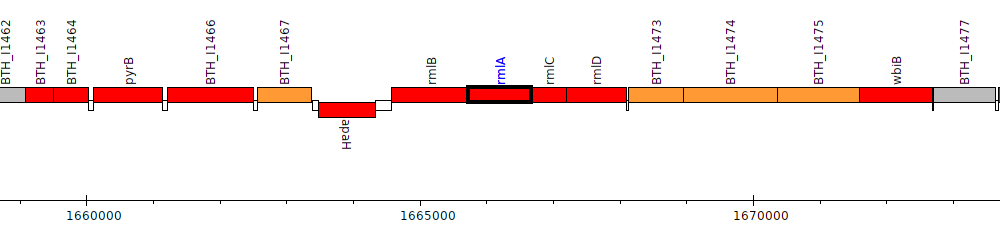
Burkholderia thailandensis E264 ATCC 700388, BTH_I1470 (rmlA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016779 | nucleotidyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00483
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008879 | glucose-1-phosphate thymidylyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01207
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0045226 | extracellular polysaccharide biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01207
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009058 | biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF00483
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | dTDP-<i>N</i>-acetylviosamine biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | dTDP-D-ravidosamine and dTDP-4-acetyl-D-ravidosamine biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00523 | Polyketide sugar unit biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | dTDP-D-olivose, dTDP-D-oliose and dTDP-D-mycarose biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | dTDP-α-D-mycaminose biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | dTDP-L-daunosamine biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bte01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00525 | Acarbose and validamycin biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bte01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte00523 | Polyketide sugar unit biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | dTDP-6-deoxy-α-D-allose biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | dTDP-β-L-4-<i>epi</i>-vancosamine biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00521 | Streptomycin biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bte01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte00521 | Streptomycin biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | dTDP-3-acetamido-α-D-fucose biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | dTDP-D-desosamine biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | dTDP-D-β-fucofuranose biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | dTDP-4-<i>O</i>-demethyl-β-L-noviose biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | dTDP-D-forosamine biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | dTDP-β-L-digitoxose biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | dTDP-L-mycarose biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | dTDP-L-megosamine biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | dTDP-L-olivose biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | dTDP-<i>N</i>-acetylthomosamine biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | dTDP-3-acetamido-3,6-dideoxy-α-D-glucose biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00483 | Nucleotidyl transferase | IPR005835 | Nucleotidyl transferase domain | 19 | 255 | 4.4E-73 |
| CDD | cd02538 | G1P_TT_short | IPR005907 | Glucose-1-phosphate thymidylyltransferase, short form | 18 | 257 | 5.15297E-174 |
| SUPERFAMILY | SSF53448 | IPR029044 | Nucleotide-diphospho-sugar transferases | 17 | 303 | 4.73E-95 | |
| TIGRFAM | TIGR01207 | rmlA: glucose-1-phosphate thymidylyltransferase | IPR005907 | Glucose-1-phosphate thymidylyltransferase, short form | 19 | 303 | 1.1E-160 |
| Gene3D | G3DSA:3.90.550.10 | IPR029044 | Nucleotide-diphospho-sugar transferases | 9 | 312 | 5.0E-93 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.