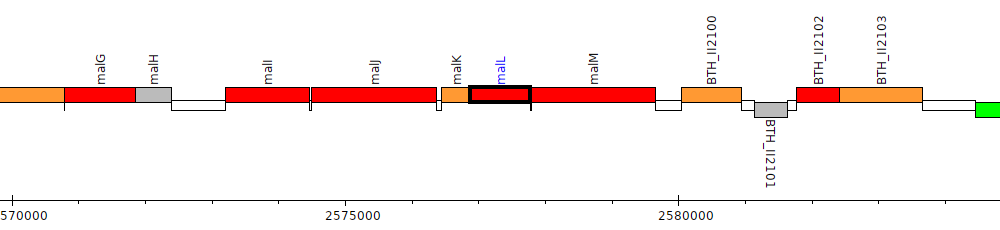
Burkholderia thailandensis E264 ATCC 700388, BTH_II2098 (malL)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004314 | [acyl-carrier-protein] S-malonyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00128
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016740 | transferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.366.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | mupirocin biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | fatty acid biosynthesis (plant mitochondria) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | pederin biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | fatty acid biosynthesis initiation I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00061 | Fatty acid biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | bryostatin biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bte01212 | Fatty acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte00061 | Fatty acid biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | octanoyl-[acyl-carrier protein] biosynthesis (mitochondria, yeast) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF52151 | IPR016035 | Acyl transferase/acyl hydrolase/lysophospholipase | 2 | 284 | 1.2E-59 | |
| Gene3D | G3DSA:3.40.366.10 | IPR001227 | Acyl transferase domain superfamily | 5 | 274 | 9.4E-85 | |
| SUPERFAMILY | SSF55048 | IPR016036 | Malonyl-CoA ACP transacylase, ACP-binding | 123 | 191 | 3.92E-11 | |
| Pfam | PF00698 | Acyl transferase domain | IPR014043 | Acyl transferase | 5 | 285 | 2.4E-33 |
| Gene3D | G3DSA:3.30.70.250 | 123 | 190 | 9.4E-85 | |||
| TIGRFAM | TIGR00128 | fabD: malonyl CoA-acyl carrier protein transacylase | IPR004410 | Malonyl CoA-acyl carrier protein transacylase, FabD-type | 2 | 285 | 3.5E-83 |
| SMART | SM00827 | Acyl transferase domain in polyketide synthase (PKS) enzymes. | IPR020801 | Polyketide synthase, acyl transferase domain | 5 | 287 | 1.1E-27 |
| PIRSF | PIRSF000446 | IPR024925 | Malonyl CoA-acyl carrier protein transacylase | 1 | 296 | 2.5E-72 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.