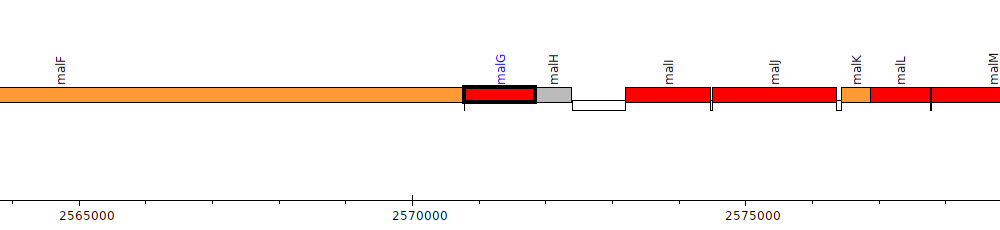
Burkholderia thailandensis E264 ATCC 700388, BTH_II2094 (malG)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009082 | branched-chain amino acid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF01450
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004455 | ketol-acid reductoisomerase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01450
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:SSF48179
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bte01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | L-isoleucine biosynthesis III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | pyruvate fermentation to isobutanol (engineered) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00770 | Pantothenate and CoA biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00290 | Valine, leucine and isoleucine biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bte00290 | Valine, leucine and isoleucine biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00270 | Cysteine and methionine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bte01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte00770 | Pantothenate and CoA biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | <i>S</i>-adenosyl-L-methionine cycle II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bte01210 | 2-Oxocarboxylic acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF48179 | IPR008927 | 6-phosphogluconate dehydrogenase-like, C-terminal domain superfamily | 184 | 327 | 9.86E-16 | |
| Pfam | PF07991 | Acetohydroxy acid isomeroreductase, NADPH-binding domain | IPR013116 | Ketol-acid reductoisomerase, N-terminal | 16 | 178 | 1.1E-46 |
| Pfam | PF01450 | Acetohydroxy acid isomeroreductase, catalytic domain | IPR000506 | Ketol-acid reductoisomerase, C-terminal | 185 | 327 | 4.4E-23 |
| Gene3D | G3DSA:1.10.3730.40 | 183 | 335 | 1.6E-28 | |||
| SMART | SM00997 | IPR015878 | S-adenosyl-L-homocysteine hydrolase, NAD binding domain | 10 | 146 | 0.001 | |
| ProSiteProfiles | PS51851 | KARI C-terminal domain profile. | IPR000506 | Ketol-acid reductoisomerase, C-terminal | 183 | 330 | 10.517 |
| ProSiteProfiles | PS51850 | KARI N-terminal domain profile. | IPR013116 | Ketol-acid reductoisomerase, N-terminal | 1 | 182 | 40.501 |
| SUPERFAMILY | SSF51735 | IPR036291 | NAD(P)-binding domain superfamily | 5 | 180 | 4.23E-49 | |
| Gene3D | G3DSA:3.40.50.720 | 4 | 182 | 2.4E-57 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.