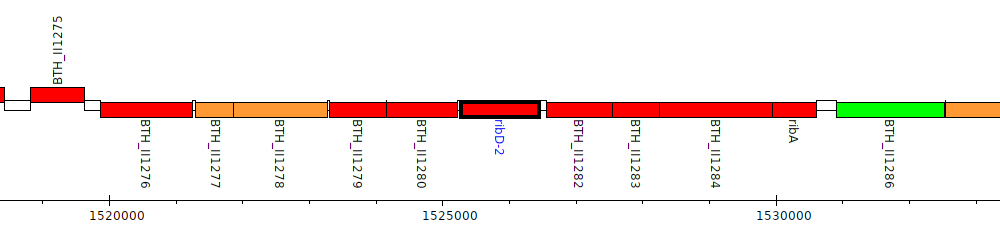
Burkholderia thailandensis E264 ATCC 700388, BTH_II1281 (ribD-2)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0008270 | zinc ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS00903
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016787 | hydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS00903
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009231 | riboflavin biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00326
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008835 | diaminohydroxyphosphoribosylaminopyrimidine deaminase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00326
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF53927
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00326
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bte01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte00740 | Riboflavin metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | toxoflavin biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00740 | Riboflavin metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.430.10 | IPR024072 | Dihydrofolate reductase-like domain superfamily | 142 | 340 | 1.8E-9 | |
| TIGRFAM | TIGR00326 | eubact_ribD: riboflavin biosynthesis protein RibD | IPR004794 | Riboflavin biosynthesis protein RibD | 12 | 198 | 1.4E-62 |
| Gene3D | G3DSA:3.40.140.10 | 3 | 140 | 9.4E-43 | |||
| ProSiteProfiles | PS51747 | Cytidine and deoxycytidylate deaminases domain profile. | IPR002125 | Cytidine and deoxycytidylate deaminase domain | 6 | 128 | 23.829 |
| CDD | cd01284 | Riboflavin_deaminase-reductase | 12 | 124 | 6.61097E-49 | ||
| Pfam | PF00383 | Cytidine and deoxycytidylate deaminase zinc-binding region | IPR002125 | Cytidine and deoxycytidylate deaminase domain | 9 | 104 | 1.7E-17 |
| PIRSF | PIRSF006769 | IPR004794 | Riboflavin biosynthesis protein RibD | 4 | 201 | 1.3E-70 | |
| SUPERFAMILY | SSF53927 | IPR016193 | Cytidine deaminase-like | 8 | 149 | 6.02E-44 | |
| PIRSF | PIRSF006769 | IPR004794 | Riboflavin biosynthesis protein RibD | 253 | 374 | 1.3E-6 | |
| ProSitePatterns | PS00903 | Cytidine and deoxycytidylate deaminases zinc-binding region signature. | IPR016192 | APOBEC/CMP deaminase, zinc-binding | 55 | 93 | - |
| SUPERFAMILY | SSF53597 | IPR024072 | Dihydrofolate reductase-like domain superfamily | 150 | 373 | 4.77E-9 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.