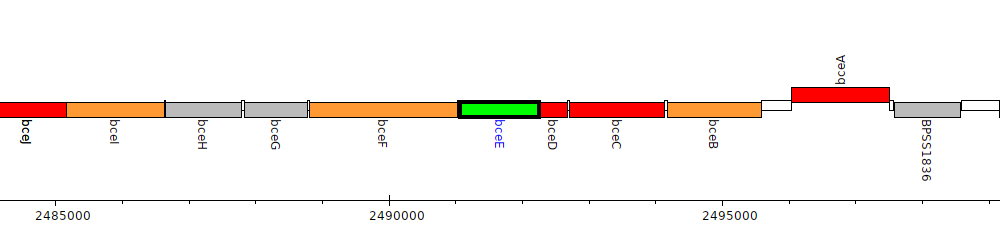
Burkholderia pseudomallei K96243, BPSS1831 (bceE)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0015159 | polysaccharide transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF02563
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0015774 | polysaccharide transport |
Inferred from Sequence Model
Term mapped from: InterPro:PF02563
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PF02563
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.30.1950.10 | 85 | 190 | 6.9E-99 | |||
| ProSiteProfiles | PS51257 | Prokaryotic membrane lipoprotein lipid attachment site profile. | 1 | 22 | 6.0 | ||
| Pfam | PF10531 | SLBB domain | IPR019554 | Soluble ligand binding domain | 193 | 241 | 6.4E-8 |
| Pfam | PF10531 | SLBB domain | IPR019554 | Soluble ligand binding domain | 277 | 329 | 6.6E-6 |
| Gene3D | G3DSA:3.10.560.10 | 51 | 361 | 6.9E-99 | |||
| Pfam | PF02563 | Polysaccharide biosynthesis/export protein | IPR003715 | Polysaccharide export protein | 79 | 186 | 2.2E-27 |
| Gene3D | G3DSA:3.10.560.10 | 62 | 275 | 6.9E-99 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.