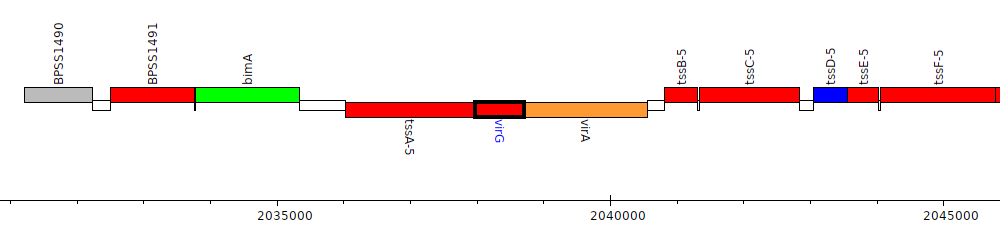
Burkholderia pseudomallei K96243, BPSS1494 (virG)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0033103 | protein secretion by the type VI secretion system | Traceable Author Statement | ECO:0000304 traceable author statement used in manual assertion |
17660433 | Reviewed by curator |
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:cd00383
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0000160 | phosphorelay signal transduction system |
Inferred from Sequence Model
Term mapped from: InterPro:cd00383
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd00383
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:cd00383
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd00383
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0000160 | phosphorelay signal transduction system |
Inferred from Sequence Model
Term mapped from: InterPro:cd00383
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF52172 | IPR011006 | CheY-like superfamily | 6 | 131 | 8.11E-33 | |
| ProSiteProfiles | PS51755 | OmpR/PhoB-type DNA-binding domain profile. | IPR001867 | OmpR/PhoB-type DNA-binding domain | 138 | 237 | 35.858 |
| SMART | SM00862 | IPR001867 | OmpR/PhoB-type DNA-binding domain | 160 | 235 | 1.3E-22 | |
| CDD | cd00383 | trans_reg_C | IPR001867 | OmpR/PhoB-type DNA-binding domain | 146 | 235 | 6.58373E-28 |
| SUPERFAMILY | SSF46894 | IPR016032 | Signal transduction response regulator, C-terminal effector | 141 | 235 | 1.93E-24 | |
| Gene3D | G3DSA:3.40.50.2300 | 1 | 127 | 1.6E-30 | |||
| SMART | SM00448 | IPR001789 | Signal transduction response regulator, receiver domain | 8 | 118 | 1.0E-29 | |
| Pfam | PF00072 | Response regulator receiver domain | IPR001789 | Signal transduction response regulator, receiver domain | 10 | 118 | 6.8E-20 |
| ProSiteProfiles | PS50110 | Response regulatory domain profile. | IPR001789 | Signal transduction response regulator, receiver domain | 9 | 122 | 33.323 |
| Pfam | PF00486 | Transcriptional regulatory protein, C terminal | IPR001867 | OmpR/PhoB-type DNA-binding domain | 161 | 235 | 2.3E-20 |
| CDD | cd00156 | REC | IPR001789 | Signal transduction response regulator, receiver domain | 11 | 121 | 5.16319E-30 |
| Gene3D | G3DSA:1.10.10.10 | IPR036388 | Winged helix-like DNA-binding domain superfamily | 135 | 241 | 3.4E-31 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.