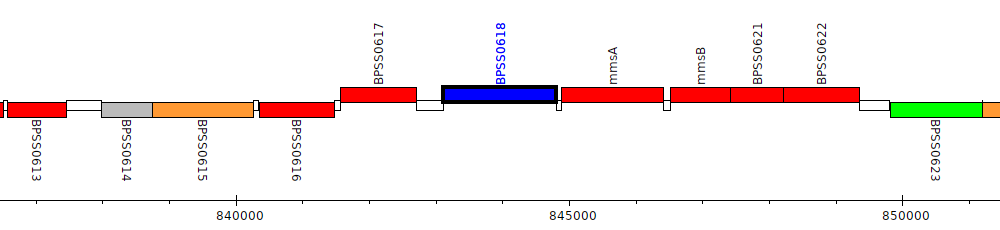
Burkholderia pseudomallei K96243, BPSS0618
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00501
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps00640 | Propanoate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00620 | Pyruvate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00680 | Methane metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00010 | Glycolysis / Gluconeogenesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01200 | Carbon metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00501 | AMP-binding enzyme | IPR000873 | AMP-dependent synthetase/ligase | 46 | 446 | 6.2E-74 |
| Gene3D | G3DSA:3.40.50.12780 | IPR042099 | AMP-dependent synthetase-like superfamily | 12 | 440 | 7.3E-113 | |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 544 | 567 | - | ||
| CDD | cd05974 | MACS_like_1 | 67 | 533 | 0.0 | ||
| ProSitePatterns | PS00455 | Putative AMP-binding domain signature. | IPR020845 | AMP-binding, conserved site | 198 | 209 | - |
| Pfam | PF13193 | AMP-binding enzyme C-terminal domain | IPR025110 | AMP-binding enzyme, C-terminal domain | 455 | 532 | 3.3E-16 |
| SUPERFAMILY | SSF56801 | 13 | 544 | 2.09E-130 | |||
| Gene3D | G3DSA:3.30.300.310 | 442 | 546 | 3.0E-20 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.