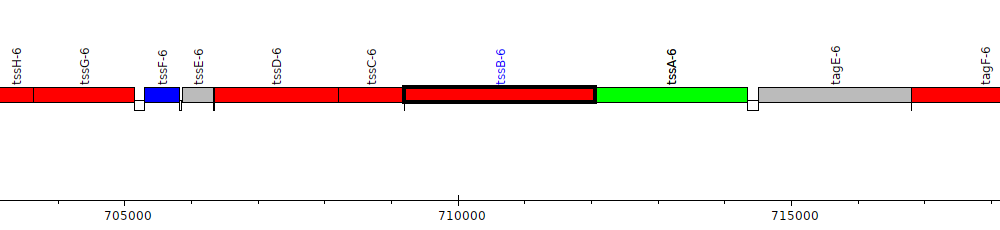
Burkholderia pseudomallei K96243, BPSS0522 (tssB-6)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0033103 | protein secretion by the type VI secretion system | Traceable Author Statement | ECO:0000304 traceable author statement used in manual assertion |
17660433 | Reviewed by curator |
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00004
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0019538 | protein metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:SSF81923
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps03070 | Bacterial secretion system | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 547 | 838 | 6.0E-59 | |
| Pfam | PF07724 | AAA domain (Cdc48 subfamily) | IPR003959 | ATPase, AAA-type, core | 597 | 763 | 1.3E-33 |
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 184 | 445 | 3.91E-60 | |
| SMART | SM00382 | IPR003593 | AAA+ ATPase domain | 598 | 742 | 3.6E-9 | |
| Gene3D | G3DSA:3.40.50.300 | 373 | 545 | 4.9E-32 | |||
| PRINTS | PR00300 | ATP-dependent Clp protease ATP-binding subunit signature | IPR001270 | ClpA/B family | 676 | 694 | 5.3E-26 |
| Gene3D | G3DSA:1.10.1780.10 | IPR036628 | Clp, N-terminal domain superfamily | 1 | 159 | 3.0E-23 | |
| Pfam | PF00004 | ATPase family associated with various cellular activities (AAA) | IPR003959 | ATPase, AAA-type, core | 228 | 333 | 5.7E-8 |
| PRINTS | PR00300 | ATP-dependent Clp protease ATP-binding subunit signature | IPR001270 | ClpA/B family | 709 | 723 | 5.3E-26 |
| SUPERFAMILY | SSF81923 | IPR036628 | Clp, N-terminal domain superfamily | 14 | 124 | 2.88E-11 | |
| SMART | SM00382 | IPR003593 | AAA+ ATPase domain | 223 | 368 | 2.9E-8 | |
| CDD | cd00009 | AAA | 206 | 366 | 1.39742E-15 | ||
| SMART | SM01086 | IPR019489 | Clp ATPase, C-terminal | 769 | 859 | 3.2E-4 | |
| Gene3D | G3DSA:3.40.50.300 | 551 | 767 | 7.9E-62 | |||
| Gene3D | G3DSA:3.40.50.300 | 183 | 372 | 6.2E-58 | |||
| PRINTS | PR00300 | ATP-dependent Clp protease ATP-binding subunit signature | IPR001270 | ClpA/B family | 647 | 665 | 5.3E-26 |
| ProSitePatterns | PS00870 | Chaperonins clpA/B signature 1. | IPR018368 | ClpA/B, conserved site 1 | 319 | 331 | - |
| PRINTS | PR00300 | ATP-dependent Clp protease ATP-binding subunit signature | IPR001270 | ClpA/B family | 602 | 620 | 5.3E-26 |
| CDD | cd00009 | AAA | 595 | 727 | 5.60885E-15 | ||
| Coils | Coil | 956 | 956 | - | |||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 898 | 956 | - | ||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 922 | 937 | - | ||
| Pfam | PF10431 | C-terminal, D2-small domain, of ClpB protein | IPR019489 | Clp ATPase, C-terminal | 769 | 842 | 2.4E-6 |
| Pfam | PF02861 | Clp amino terminal domain, pathogenicity island component | IPR004176 | Clp, N-terminal | 23 | 72 | 4.9E-4 |
| TIGRFAM | TIGR03345 | VI_ClpV1: type VI secretion ATPase, ClpV1 family | IPR017729 | Type VI secretion system, ATPase ClpV1 | 12 | 852 | 6.9E-281 |
| Pfam | PF17871 | AAA lid domain | IPR041546 | ClpA/ClpB, AAA lid domain | 368 | 447 | 1.8E-25 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.