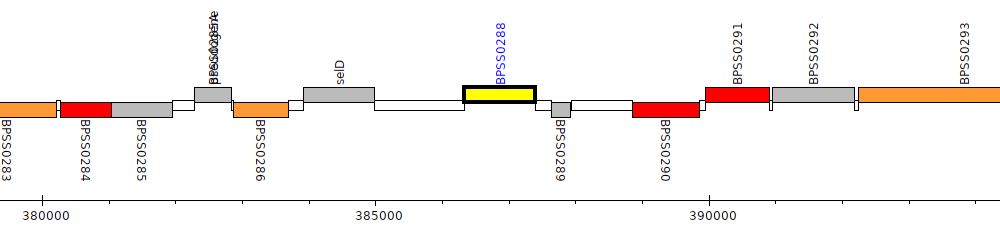
Burkholderia pseudomallei K96243, BPSS0288
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.50.1580
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009116 | nucleoside metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.50.1580
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055085 | transmembrane transport |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF013171
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PIRSF | PIRSF013171 | IPR009486 | Purine nucleoside permease | 2 | 353 | 6.9E-151 | |
| ProSiteProfiles | PS51257 | Prokaryotic membrane lipoprotein lipid attachment site profile. | 1 | 17 | 5.0 | ||
| CDD | cd17766 | futalosine_nucleosidase_MqnB | 81 | 138 | 0.0056363 | ||
| Pfam | PF06516 | Purine nucleoside permease (NUP) | IPR009486 | Purine nucleoside permease | 41 | 351 | 2.5E-128 |
| SUPERFAMILY | SSF53167 | IPR035994 | Nucleoside phosphorylase superfamily | 42 | 184 | 2.05E-5 | |
| Gene3D | G3DSA:3.40.50.1580 | IPR035994 | Nucleoside phosphorylase superfamily | 31 | 300 | 1.3E-8 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.