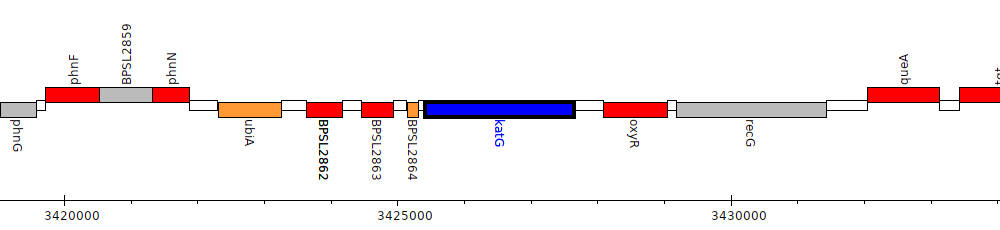
Burkholderia pseudomallei K96243, BPSL2865 (katG)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004601 | peroxidase activity | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
12832453 | Reviewed by curator |
| Molecular Function | GO:0004096 | catalase activity | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
12832453 | Reviewed by curator |
| Molecular Function | GO:0004096 | catalase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00198
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006979 | response to oxidative stress |
Inferred from Sequence Model
Term mapped from: InterPro:SSF48113
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004601 | peroxidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF48113
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:SSF48113
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0020037 | heme binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF48113
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00940 | Phenylpropanoid biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | methanol oxidation to formaldehyde IV | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00983 | Drug metabolism - other enzymes | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00380 | Tryptophan metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00360 | Phenylalanine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps00380 | Tryptophan metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00360 | Phenylalanine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| MobiDBLite | mobidb-lite | consensus disorder prediction | 1 | 26 | - | ||
| SUPERFAMILY | SSF48113 | IPR010255 | Haem peroxidase superfamily | 35 | 442 | 6.34E-155 | |
| PRINTS | PR00460 | Bacterial haem catalase-peroxidase signature | IPR000763 | Catalase-peroxidase haem | 65 | 78 | 9.4E-57 |
| Gene3D | G3DSA:1.10.520.10 | 444 | 747 | 6.9E-155 | |||
| PRINTS | PR00458 | Haem peroxidase superfamily signature | IPR002016 | Haem peroxidase | 330 | 345 | 9.7E-19 |
| CDD | cd00649 | catalase_peroxidase_1 | 34 | 444 | 0.0 | ||
| ProSitePatterns | PS00436 | Peroxidases active site signature. | IPR019794 | Peroxidase, active site | 103 | 114 | - |
| Gene3D | G3DSA:1.10.520.10 | 33 | 442 | 1.0E-225 | |||
| PRINTS | PR00458 | Haem peroxidase superfamily signature | IPR002016 | Haem peroxidase | 271 | 286 | 9.7E-19 |
| Gene3D | G3DSA:1.10.420.10 | 600 | 723 | 6.9E-155 | |||
| ProSitePatterns | PS00435 | Peroxidases proximal heme-ligand signature. | IPR019793 | Peroxidases heam-ligand binding site | 271 | 281 | - |
| ProSiteProfiles | PS50873 | Plant heme peroxidase family profile. | IPR002016 | Haem peroxidase | 145 | 443 | 10.902 |
| PRINTS | PR00460 | Bacterial haem catalase-peroxidase signature | IPR000763 | Catalase-peroxidase haem | 80 | 105 | 9.4E-57 |
| CDD | cd08200 | catalase_peroxidase_2 | 448 | 744 | 0.0 | ||
| PRINTS | PR00458 | Haem peroxidase superfamily signature | IPR002016 | Haem peroxidase | 181 | 193 | 9.7E-19 |
| Pfam | PF00141 | Peroxidase | IPR002016 | Haem peroxidase | 90 | 407 | 3.0E-48 |
| Pfam | PF00141 | Peroxidase | IPR002016 | Haem peroxidase | 414 | 720 | 1.1E-37 |
| SUPERFAMILY | SSF48113 | IPR010255 | Haem peroxidase superfamily | 442 | 747 | 2.06E-115 | |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 8 | 24 | - | ||
| PRINTS | PR00458 | Haem peroxidase superfamily signature | IPR002016 | Haem peroxidase | 103 | 117 | 9.7E-19 |
| Gene3D | G3DSA:1.10.420.20 | 203 | 409 | 1.0E-225 | |||
| PRINTS | PR00460 | Bacterial haem catalase-peroxidase signature | IPR000763 | Catalase-peroxidase haem | 492 | 518 | 9.4E-57 |
| PRINTS | PR00460 | Bacterial haem catalase-peroxidase signature | IPR000763 | Catalase-peroxidase haem | 39 | 61 | 9.4E-57 |
| PRINTS | PR00460 | Bacterial haem catalase-peroxidase signature | IPR000763 | Catalase-peroxidase haem | 279 | 302 | 9.4E-57 |
| Hamap | MF_01961 | Catalase-peroxidase [katG]. | IPR000763 | Catalase-peroxidase haem | 22 | 748 | 42.63 |
| TIGRFAM | TIGR00198 | cat_per_HPI: catalase/peroxidase HPI | IPR000763 | Catalase-peroxidase haem | 26 | 747 | 0.0 |
| PRINTS | PR00458 | Haem peroxidase superfamily signature | IPR002016 | Haem peroxidase | 163 | 180 | 9.7E-19 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.