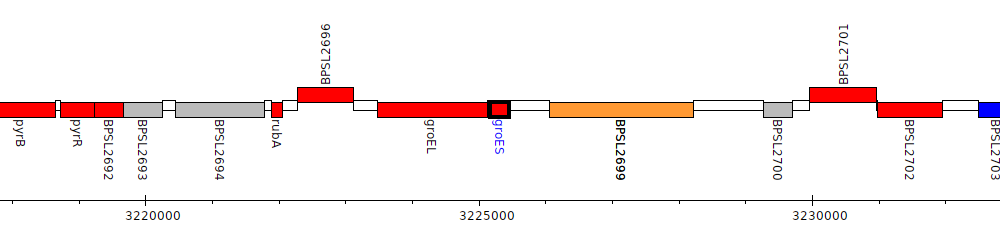
Burkholderia pseudomallei K96243, BPSL2698 (groES)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006457 | protein folding |
Inferred from Sequence Model
Term mapped from: InterPro:PS00681
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS00681
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00883 | IPR020818 | GroES chaperonin family | 2 | 94 | 2.9E-57 | |
| PRINTS | PR00297 | 10kDa chaperonin signature | IPR020818 | GroES chaperonin family | 3 | 18 | 1.4E-33 |
| PRINTS | PR00297 | 10kDa chaperonin signature | IPR020818 | GroES chaperonin family | 81 | 94 | 1.4E-33 |
| ProSitePatterns | PS00681 | Chaperonins cpn10 signature. | IPR018369 | Chaperonin GroES, conserved site | 3 | 27 | - |
| Pfam | PF00166 | Chaperonin 10 Kd subunit | IPR020818 | GroES chaperonin family | 2 | 94 | 1.4E-35 |
| CDD | cd00320 | cpn10 | IPR020818 | GroES chaperonin family | 2 | 94 | 6.36087E-43 |
| SUPERFAMILY | SSF50129 | IPR011032 | GroES-like superfamily | 1 | 95 | 2.47E-34 | |
| PRINTS | PR00297 | 10kDa chaperonin signature | IPR020818 | GroES chaperonin family | 25 | 46 | 1.4E-33 |
| Hamap | MF_00580 | 10 kDa chaperonin [groS]. | IPR020818 | GroES chaperonin family | 2 | 95 | 26.725 |
| Gene3D | G3DSA:2.30.33.40 | IPR037124 | GroES chaperonin superfamily | 1 | 97 | 1.5E-38 | |
| PRINTS | PR00297 | 10kDa chaperonin signature | IPR020818 | GroES chaperonin family | 60 | 72 | 1.4E-33 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.