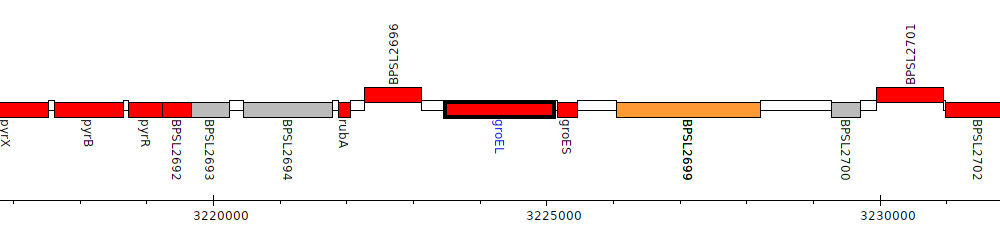
Burkholderia pseudomallei K96243, BPSL2697 (groEL)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00118
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006457 | protein folding |
Inferred from Sequence Model
Term mapped from: InterPro:PS00296
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005737 | cytoplasm |
Inferred from Sequence Model
Term mapped from: InterPro:PS00296
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0042026 | protein refolding |
Inferred from Sequence Model
Term mapped from: InterPro:cd03344
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps03018 | RNA degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd03344 | GroEL | IPR001844 | Chaperonin Cpn60 | 4 | 523 | 0.0 |
| ProSitePatterns | PS00296 | Chaperonins cpn60 signature. | IPR018370 | Chaperonin Cpn60, conserved site | 405 | 416 | - |
| PRINTS | PR00298 | 60kDa chaperonin signature | IPR001844 | Chaperonin Cpn60 | 268 | 291 | 8.2E-75 |
| Gene3D | G3DSA:1.10.560.10 | IPR027413 | GroEL-like equatorial domain superfamily | 8 | 521 | 8.7E-238 | |
| SUPERFAMILY | SSF48592 | IPR027413 | GroEL-like equatorial domain superfamily | 7 | 521 | 3.97E-78 | |
| SUPERFAMILY | SSF54849 | IPR027410 | TCP-1-like chaperonin intermediate domain superfamily | 137 | 202 | 1.65E-24 | |
| Gene3D | G3DSA:3.50.7.10 | IPR027409 | GroEL-like apical domain superfamily | 192 | 372 | 8.7E-238 | |
| PRINTS | PR00298 | 60kDa chaperonin signature | IPR001844 | Chaperonin Cpn60 | 350 | 375 | 8.2E-75 |
| Coils | Coil | 339 | 366 | - | |||
| PRINTS | PR00298 | 60kDa chaperonin signature | IPR001844 | Chaperonin Cpn60 | 398 | 419 | 8.2E-75 |
| SUPERFAMILY | SSF52029 | IPR027409 | GroEL-like apical domain superfamily | 185 | 376 | 1.14E-82 | |
| Hamap | MF_00600 | 60 kDa chaperonin [groL]. | IPR001844 | Chaperonin Cpn60 | 2 | 542 | 41.708 |
| Pfam | PF00118 | TCP-1/cpn60 chaperonin family | IPR002423 | Chaperonin Cpn60/TCP-1 family | 23 | 522 | 9.1E-100 |
| TIGRFAM | TIGR02348 | GroEL: chaperonin GroL | IPR001844 | Chaperonin Cpn60 | 3 | 527 | 1.2E-254 |
| PRINTS | PR00298 | 60kDa chaperonin signature | IPR001844 | Chaperonin Cpn60 | 27 | 53 | 8.2E-75 |
| PRINTS | PR00298 | 60kDa chaperonin signature | IPR001844 | Chaperonin Cpn60 | 83 | 110 | 8.2E-75 |
| Gene3D | G3DSA:3.30.260.10 | IPR027410 | TCP-1-like chaperonin intermediate domain superfamily | 136 | 410 | 8.7E-238 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.