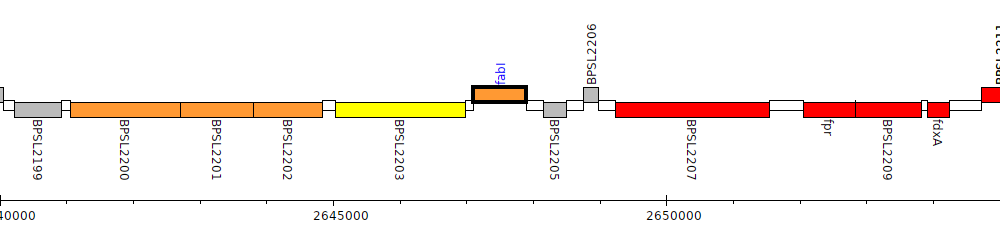
Burkholderia pseudomallei K96243, BPSL2204 (fabI)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006633 | fatty acid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000094
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000094
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004318 | enoyl-[acyl-carrier-protein] reductase (NADH) activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000094
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | mycolate biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | palmitoleate biosynthesis I (from (5Z)-dodec-5-enoate) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | superpathway of mycolate biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps00061 | Fatty acid biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | 8-amino-7-oxononanoate biosynthesis I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | gondoate biosynthesis (anaerobic) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | oleate biosynthesis IV (anaerobic) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | (5Z)-dodecenoate biosynthesis II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00061 | Fatty acid biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | palmitate biosynthesis II (bacteria and plants) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | stearate biosynthesis II (bacteria and plants) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps00780 | Biotin metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | <i>cis</i>-vaccenate biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps01212 | Fatty acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd05372 | ENR_SDR | IPR014358 | Enoyl-[acyl-carrier-protein] reductase (NADH) | 6 | 254 | 2.43378E-139 |
| Pfam | PF13561 | Enoyl-(Acyl carrier protein) reductase | 18 | 252 | 9.5E-87 | ||
| SUPERFAMILY | SSF51735 | IPR036291 | NAD(P)-binding domain superfamily | 5 | 253 | 3.25E-62 | |
| Gene3D | G3DSA:3.40.50.720 | 1 | 258 | 4.9E-56 | |||
| PIRSF | PIRSF000094 | IPR014358 | Enoyl-[acyl-carrier-protein] reductase (NADH) | 1 | 260 | 6.2E-114 | |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 215 | 235 | 6.5E-9 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 180 | 197 | 6.5E-9 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 159 | 178 | 6.5E-9 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 83 | 94 | 6.5E-9 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.