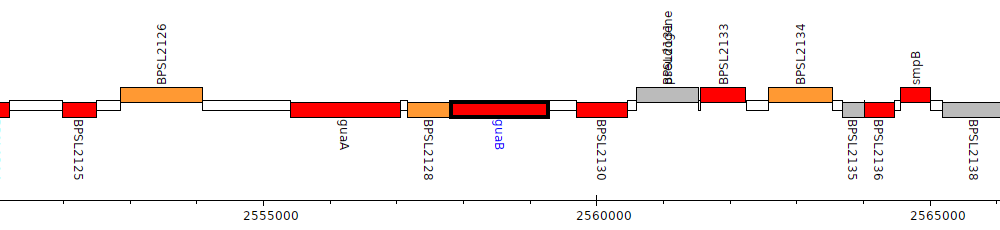
Burkholderia pseudomallei K96243, BPSL2129 (guaB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01964
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006164 | purine nucleotide biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01964
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003938 | IMP dehydrogenase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01964
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS00487
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.20.20.70
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.20.20.70
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01964
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006164 | purine nucleotide biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01964
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS00487
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003938 | IMP dehydrogenase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01964
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps00230 | Purine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00230 | Purine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | inosine 5'-phosphate degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | guanosine ribonucleotides <i>de novo</i> biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | adenosine nucleotides degradation I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00983 | Drug metabolism - other enzymes | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF51412 | 2 | 476 | 4.58E-126 | |||
| SMART | SM01240 | 6 | 472 | 0.0 | |||
| Gene3D | G3DSA:3.20.20.70 | IPR013785 | Aldolase-type TIM barrel | 1 | 485 | 9.6E-216 | |
| SMART | SM00116 | IPR000644 | CBS domain | 95 | 142 | 2.2E-11 | |
| TIGRFAM | TIGR01302 | IMP_dehydrog: inosine-5'-monophosphate dehydrogenase | IPR005990 | Inosine-5'-monophosphate dehydrogenase | 7 | 454 | 1.2E-190 |
| Hamap | MF_01964 | Inosine-5'-monophosphate dehydrogenase [guaB]. | IPR005990 | Inosine-5'-monophosphate dehydrogenase | 7 | 485 | 85.218 |
| Pfam | PF00571 | CBS domain | IPR000644 | CBS domain | 94 | 138 | 1.3E-9 |
| ProSitePatterns | PS00487 | IMP dehydrogenase / GMP reductase signature. | IPR015875 | IMP dehydrogenase / GMP reductase, conserved site | 293 | 305 | - |
| PIRSF | PIRSF000130 | IPR005990 | Inosine-5'-monophosphate dehydrogenase | 1 | 486 | 2.7E-207 | |
| CDD | cd04601 | CBS_pair_IMPDH | 93 | 201 | 1.23773E-51 | ||
| SMART | SM00116 | IPR000644 | CBS domain | 156 | 204 | 7.5E-8 | |
| Pfam | PF00478 | IMP dehydrogenase / GMP reductase domain | IPR001093 | IMP dehydrogenase/GMP reductase | 7 | 472 | 6.4E-165 |
| CDD | cd00381 | IMPDH | IPR001093 | IMP dehydrogenase/GMP reductase | 7 | 461 | 3.86149E-166 |
| Pfam | PF00571 | CBS domain | IPR000644 | CBS domain | 147 | 200 | 1.1E-6 |
| SUPERFAMILY | SSF54631 | 93 | 201 | 3.78E-28 | |||
| ProSiteProfiles | PS51371 | CBS domain profile. | IPR000644 | CBS domain | 92 | 149 | 12.402 |
| ProSiteProfiles | PS51371 | CBS domain profile. | IPR000644 | CBS domain | 151 | 212 | 12.1 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.