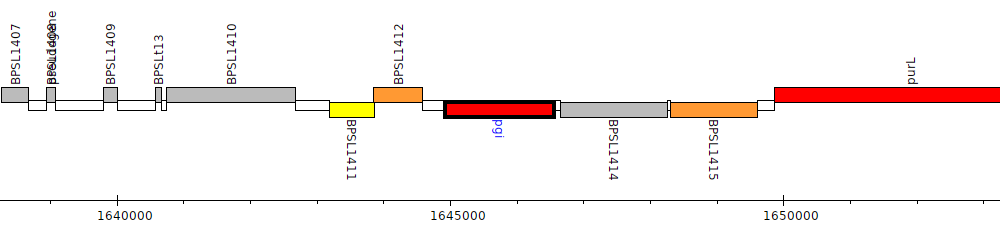
Burkholderia pseudomallei K96243, BPSL1413 (pgi)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006094 | gluconeogenesis |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00473
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004347 | glucose-6-phosphate isomerase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00473
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006096 | glycolytic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00473
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | sucrose biosynthesis III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00520 | Amino sugar and nucleotide sugar metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | D-sorbitol biosynthesis I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | UDP-<i>N</i>-acetyl-D-galactosamine biosynthesis II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | gluconeogenesis II (<i>Methanobacterium thermoautotrophicum</i>) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | starch biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | 1,5-anhydrofructose degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00010 | Glycolysis / Gluconeogenesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | sucrose degradation IV (sucrose phosphorylase) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | chitin biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00030 | Pentose phosphate pathway | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00500 | Starch and sucrose metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00010 | Glycolysis / Gluconeogenesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | sucrose degradation II (sucrose synthase) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | sucrose degradation III (sucrose invertase) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps00500 | Starch and sucrose metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00030 | Pentose phosphate pathway | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | 1,3-propanediol biosynthesis (engineered) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps00520 | Amino sugar and nucleotide sugar metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01200 | Carbon metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | sucrose biosynthesis II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | GDP-mannose biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | UDP-<i>N</i>-acetyl-D-galactosamine biosynthesis III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00662 | Glucose-6-phosphate isomerase signature | IPR001672 | Phosphoglucose isomerase (PGI) | 333 | 354 | 2.4E-52 |
| PRINTS | PR00662 | Glucose-6-phosphate isomerase signature | IPR001672 | Phosphoglucose isomerase (PGI) | 454 | 472 | 2.4E-52 |
| Gene3D | G3DSA:1.10.1390.10 | IPR023096 | Phosphoglucose isomerase, C-terminal | 496 | 538 | 6.3E-15 | |
| PRINTS | PR00662 | Glucose-6-phosphate isomerase signature | IPR001672 | Phosphoglucose isomerase (PGI) | 144 | 163 | 2.4E-52 |
| Gene3D | G3DSA:3.40.50.10490 | 98 | 286 | 2.4E-212 | |||
| ProSiteProfiles | PS51463 | Glucose-6-phosphate isomerase family profile. | IPR001672 | Phosphoglucose isomerase (PGI) | 6 | 537 | 143.906 |
| PRINTS | PR00662 | Glucose-6-phosphate isomerase signature | IPR001672 | Phosphoglucose isomerase (PGI) | 486 | 499 | 2.4E-52 |
| Hamap | MF_00473 | Glucose-6-phosphate isomerase [pgi]. | IPR001672 | Phosphoglucose isomerase (PGI) | 46 | 513 | 40.103 |
| Gene3D | G3DSA:3.40.50.10490 | 9 | 495 | 2.4E-212 | |||
| PRINTS | PR00662 | Glucose-6-phosphate isomerase signature | IPR001672 | Phosphoglucose isomerase (PGI) | 472 | 486 | 2.4E-52 |
| ProSitePatterns | PS00174 | Phosphoglucose isomerase signature 2. | IPR018189 | Phosphoglucose isomerase, conserved site | 486 | 503 | - |
| Pfam | PF00342 | Phosphoglucose isomerase | IPR001672 | Phosphoglucose isomerase (PGI) | 52 | 531 | 4.8E-196 |
| CDD | cd05015 | SIS_PGI_1 | IPR035476 | Phosphoglucose isomerase, SIS domain 1 | 116 | 279 | 3.08572E-72 |
| PRINTS | PR00662 | Glucose-6-phosphate isomerase signature | IPR001672 | Phosphoglucose isomerase (PGI) | 256 | 274 | 2.4E-52 |
| ProSitePatterns | PS00765 | Phosphoglucose isomerase signature 1. | IPR018189 | Phosphoglucose isomerase, conserved site | 260 | 273 | - |
| CDD | cd05016 | SIS_PGI_2 | IPR035482 | Phosphoglucose isomerase, SIS domain 2 | 326 | 507 | 5.94812E-76 |
| SUPERFAMILY | SSF53697 | 2 | 536 | 8.06E-198 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.