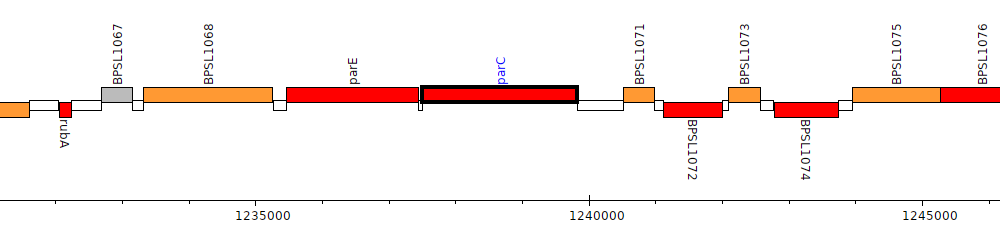
Burkholderia pseudomallei K96243, BPSL1070 (parC)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003918 | DNA topoisomerase type II (ATP-hydrolyzing) activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF56719
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005694 | chromosome |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00936
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006265 | DNA topological change |
Inferred from Sequence Model
Term mapped from: InterPro:cd00187
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006259 | DNA metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.90.199.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF56719
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003918 | DNA topoisomerase type II (ATP-hydrolyzing) activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF56719
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006265 | DNA topological change |
Inferred from Sequence Model
Term mapped from: InterPro:cd00187
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF56719
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd00187
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006259 | DNA metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.90.199.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005694 | chromosome |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00936
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd00187
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| TIGRFAM | TIGR01062 | parC_Gneg: DNA topoisomerase IV, A subunit | 20 | 756 | 1.5E-234 | ||
| SMART | SM00434 | IPR002205 | DNA topoisomerase, type IIA, subunit A/C-terminal | 20 | 492 | 4.6E-221 | |
| CDD | cd00187 | TOP4c | IPR002205 | DNA topoisomerase, type IIA, subunit A/C-terminal | 40 | 501 | 0.0 |
| Gene3D | G3DSA:3.90.199.10 | IPR013758 | DNA topoisomerase, type IIA, subunit A/ C-terminal, alpha-beta | 41 | 514 | 2.4E-188 | |
| SUPERFAMILY | SSF56719 | IPR013760 | DNA topoisomerase, type IIA-like domain superfamily | 40 | 514 | 3.53E-150 | |
| Coils | Coil | 496 | 516 | - | |||
| Gene3D | G3DSA:3.30.1360.40 | 246 | 362 | 2.4E-188 | |||
| Coils | Coil | 457 | 491 | - | |||
| Gene3D | G3DSA:1.10.268.10 | IPR013757 | Type IIA DNA topoisomerase subunit A, alpha-helical domain superfamily | 400 | 485 | 2.4E-188 | |
| Gene3D | G3DSA:2.120.10.90 | IPR035516 | DNA gyrase/topoisomerase IV, subunit A, C-terminal | 527 | 756 | 1.0E-18 | |
| Hamap | MF_00936 | DNA topoisomerase 4 subunit A [parC]. | IPR005742 | DNA topoisomerase IV, subunit A, Gram-negative | 13 | 768 | 33.259 |
| Pfam | PF00521 | DNA gyrase/topoisomerase IV, subunit A | IPR002205 | DNA topoisomerase, type IIA, subunit A/C-terminal | 41 | 498 | 5.6E-132 |
| SUPERFAMILY | SSF101904 | IPR035516 | DNA gyrase/topoisomerase IV, subunit A, C-terminal | 529 | 758 | 2.88E-46 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.