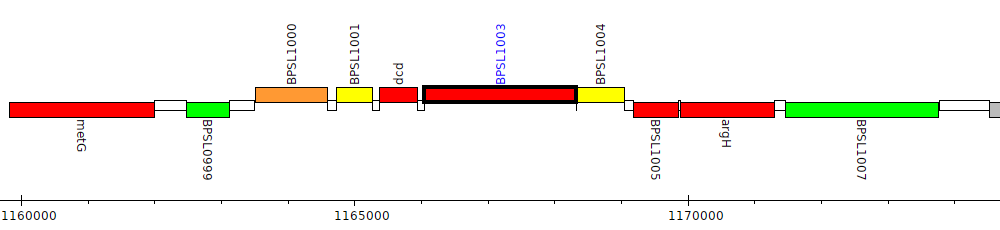
Burkholderia pseudomallei K96243, BPSL1003
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016831 | carboxy-lyase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF009393
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006520 | cellular amino acid metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF009393
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF55904
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005737 | cytoplasm |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF009393
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00330 | Arginine and proline metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Coils | Coil | 757 | 759 | - | |||
| Gene3D | G3DSA:3.90.1150.10 | IPR015422 | Pyridoxal phosphate-dependent transferase domain 1 | 468 | 638 | 1.3E-70 | |
| Gene3D | G3DSA:3.40.640.10 | IPR015421 | Pyridoxal phosphate-dependent transferase, major domain | 157 | 465 | 2.5E-97 | |
| ProSitePatterns | PS00703 | Orn/Lys/Arg decarboxylases family 1 pyridoxal-P attachment site. | IPR000310 | Orn/Lys/Arg decarboxylase, major domain | 401 | 415 | - |
| Pfam | PF01276 | Orn/Lys/Arg decarboxylase, major domain | IPR000310 | Orn/Lys/Arg decarboxylase, major domain | 157 | 592 | 8.5E-153 |
| SUPERFAMILY | SSF53383 | IPR015424 | Pyridoxal phosphate-dependent transferase | 157 | 611 | 2.76E-115 | |
| Pfam | PF03711 | Orn/Lys/Arg decarboxylase, C-terminal domain | IPR008286 | Orn/Lys/Arg decarboxylase, C-terminal | 619 | 745 | 1.2E-46 |
| Gene3D | G3DSA:3.40.50.2300 | 5 | 156 | 1.6E-20 | |||
| PIRSF | PIRSF009393 | IPR011193 | Ornithine/lysine/arginine decarboxylase | 1 | 759 | 4.0E-284 | |
| Gene3D | G3DSA:3.90.100.10 | 639 | 759 | 3.8E-39 | |||
| Pfam | PF03709 | Orn/Lys/Arg decarboxylase, N-terminal domain | IPR005308 | Orn/Lys/Arg decarboxylase, N-terminal | 20 | 151 | 1.1E-38 |
| SUPERFAMILY | SSF55904 | IPR036633 | Orn/Lys/Arg decarboxylase, C-terminal domain superfamily | 614 | 759 | 1.83E-46 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.