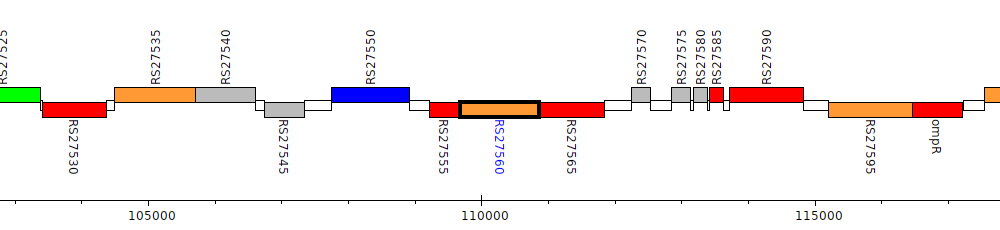
Burkholderia cenocepacia AU 1054, BCEN_RS27560
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0015703 | chromate transport |
Inferred from Sequence Model
Term mapped from: InterPro:PF02417
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0015109 | chromate transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF02417
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0015703 | chromate transport |
Inferred from Sequence Model
Term mapped from: InterPro:PF02417
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0015109 | chromate transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF02417
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PIRSF | PIRSF004810 | IPR014047 | Chromate transporter, long chain | 1 | 388 | 1.1E-88 | |
| Pfam | PF02417 | Chromate transporter | IPR003370 | Chromate transporter | 13 | 178 | 3.1E-31 |
| TIGRFAM | TIGR00937 | 2A51: chromate efflux transporter | IPR014047 | Chromate transporter, long chain | 21 | 381 | 3.0E-130 |
| Pfam | PF02417 | Chromate transporter | IPR003370 | Chromate transporter | 218 | 381 | 5.6E-33 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.