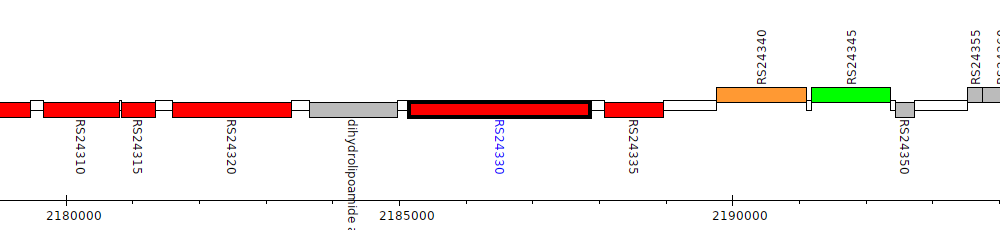
Burkholderia cenocepacia AU 1054, BCEN_RS24330
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00759
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.50.920
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00759
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00020 | Citrate cycle (TCA cycle) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | thiazole biosynthesis I (facultative anaerobic bacteria) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00620 | Pyruvate metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00730 | Thiamine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00900 | Terpenoid backbone biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00010 | Glycolysis / Gluconeogenesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | thiazole biosynthesis II (aerobic bacteria) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | methylerythritol phosphate pathway II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF52922 | IPR009014 | Transketolase C-terminal/Pyruvate-ferredoxin oxidoreductase domain II | 721 | 894 | 1.5E-52 | |
| Gene3D | G3DSA:3.40.50.970 | 68 | 482 | 4.3E-130 | |||
| TIGRFAM | TIGR03186 | AKGDH_not_PDH: alpha-ketoglutarate dehydrogenase | IPR017600 | Alpha-ketoglutarate dehydrogenase | 17 | 903 | 0.0 |
| SUPERFAMILY | SSF52518 | IPR029061 | Thiamin diphosphate-binding fold | 486 | 710 | 1.01E-91 | |
| Pfam | PF17831 | Pyruvate dehydrogenase E1 component middle domain | IPR041621 | Pyruvate dehydrogenase E1 component, middle domain | 501 | 710 | 1.8E-105 |
| Pfam | PF00456 | Transketolase, thiamine diphosphate binding domain | IPR005474 | Transketolase, N-terminal | 125 | 309 | 1.2E-7 |
| SUPERFAMILY | SSF52518 | IPR029061 | Thiamin diphosphate-binding fold | 69 | 478 | 4.91E-129 | |
| CDD | cd02017 | TPP_E1_EcPDC_like | IPR035807 | Pyruvate dehydrogenase E1 component, N-terminal | 88 | 472 | 0.0 |
| PIRSF | PIRSF000156 | IPR004660 | Pyruvate dehydrogenase E1 component | 10 | 902 | 0.0 | |
| Gene3D | G3DSA:3.40.50.920 | IPR009014 | Transketolase C-terminal/Pyruvate-ferredoxin oxidoreductase domain II | 722 | 903 | 6.7E-66 | |
| Gene3D | G3DSA:3.40.50.970 | 485 | 716 | 2.4E-100 | |||
| TIGRFAM | TIGR00759 | aceE: pyruvate dehydrogenase (acetyl-transferring), homodimeric type | IPR004660 | Pyruvate dehydrogenase E1 component | 18 | 892 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.