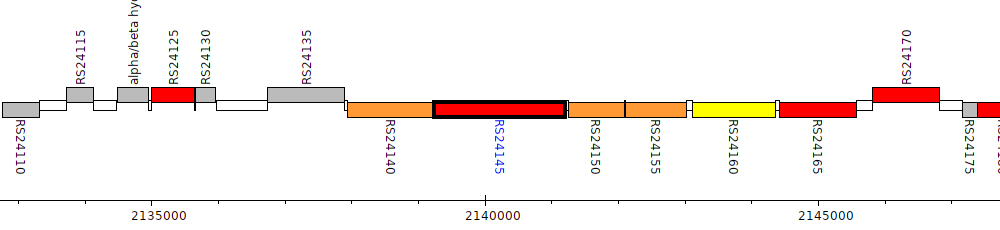
Burkholderia cenocepacia AU 1054, BCEN_RS24145
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004565 | beta-galactosidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF001084
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006012 | galactose metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF08533
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0009341 | beta-galactosidase complex |
Inferred from Sequence Model
Term mapped from: InterPro:PF02449
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0005975 | carbohydrate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF001084
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00604 | Glycosphingolipid biosynthesis - ganglio series | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | xyloglucan degradation II (exoglucanase) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00600 | Sphingolipid metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00531 | Glycosaminoglycan degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00052 | Galactose metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00511 | Other glycan degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd03143 | A4_beta-galactosidase_middle_domain | 406 | 524 | 8.22954E-29 | ||
| SUPERFAMILY | SSF51445 | IPR017853 | Glycoside hydrolase superfamily | 3 | 393 | 2.05E-124 | |
| Gene3D | G3DSA:3.40.50.880 | IPR029062 | Class I glutamine amidotransferase-like | 402 | 604 | 4.9E-58 | |
| Gene3D | G3DSA:2.60.40.1180 | IPR013780 | Glycosyl hydrolase, all-beta | 605 | 654 | 6.4E-18 | |
| Pfam | PF08532 | Beta-galactosidase trimerisation domain | IPR013738 | Beta-galactosidase trimerisation | 404 | 599 | 3.9E-34 |
| Pfam | PF08533 | Beta-galactosidase C-terminal domain | IPR013739 | Beta-galactosidase C-terminal | 607 | 653 | 9.6E-7 |
| SUPERFAMILY | SSF51011 | 602 | 654 | 7.19E-16 | |||
| SUPERFAMILY | SSF52317 | IPR029062 | Class I glutamine amidotransferase-like | 403 | 599 | 1.19E-52 | |
| Pfam | PF02449 | Beta-galactosidase | IPR013529 | Glycoside hydrolase, family 42, N-terminal | 6 | 391 | 9.2E-138 |
| Gene3D | G3DSA:3.20.20.80 | 2 | 395 | 3.2E-137 | |||
| PIRSF | PIRSF001084 | IPR003476 | Glycoside hydrolase, family 42 | 1 | 654 | 1.7E-179 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.