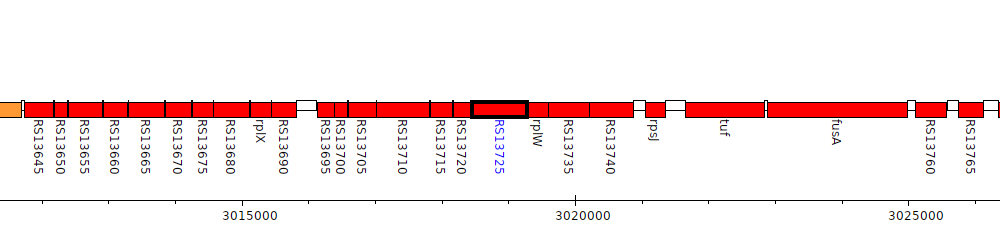
Burkholderia cenocepacia AU 1054, BCEN_RS13725
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006412 | translation |
Inferred from Sequence Model
Term mapped from: InterPro:PS00467
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016740 | transferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01171
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003723 | RNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01171
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003735 | structural constituent of ribosome |
Inferred from Sequence Model
Term mapped from: InterPro:PS00467
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0015934 | large ribosomal subunit |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01171
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005840 | ribosome |
Inferred from Sequence Model
Term mapped from: InterPro:PS00467
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:2.40.50.140 | 1 | 114 | 2.6E-47 | |||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 35 | 51 | - | ||
| Pfam | PF00181 | Ribosomal Proteins L2, RNA binding domain | IPR022666 | Ribosomal Proteins L2, RNA binding domain | 42 | 117 | 2.6E-33 |
| Gene3D | G3DSA:4.10.950.10 | IPR014726 | Ribosomal protein L2, domain 3 | 196 | 274 | 3.3E-36 | |
| SUPERFAMILY | SSF50249 | IPR012340 | Nucleic acid-binding, OB-fold | 2 | 124 | 1.1E-47 | |
| Pfam | PF03947 | Ribosomal Proteins L2, C-terminal domain | IPR022669 | Ribosomal protein L2, C-terminal | 126 | 250 | 2.6E-56 |
| SMART | SM01383 | IPR002171 | Ribosomal protein L2 | 42 | 118 | 7.8E-46 | |
| SUPERFAMILY | SSF50104 | IPR008991 | Translation protein SH3-like domain superfamily | 127 | 272 | 1.68E-60 | |
| ProSitePatterns | PS00467 | Ribosomal protein L2 signature. | IPR022671 | Ribosomal protein L2, conserved site | 218 | 229 | - |
| Gene3D | G3DSA:2.30.30.30 | IPR014722 | Ribosomal protein L2, domain 2 | 118 | 195 | 1.9E-31 | |
| PIRSF | PIRSF002158 | IPR002171 | Ribosomal protein L2 | 9 | 275 | 1.2E-101 | |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 35 | 59 | - | ||
| SMART | SM01382 | IPR022669 | Ribosomal protein L2, C-terminal | 124 | 252 | 1.7E-89 | |
| TIGRFAM | TIGR01171 | rplB_bact: ribosomal protein uL2 | IPR005880 | Ribosomal protein L2, bacterial/organellar-type | 3 | 273 | 9.3E-129 |
| Hamap | MF_01320_B | 50S ribosomal protein L2 [rplB]. | IPR005880 | Ribosomal protein L2, bacterial/organellar-type | 1 | 273 | 46.274 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 228 | 275 | - |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.