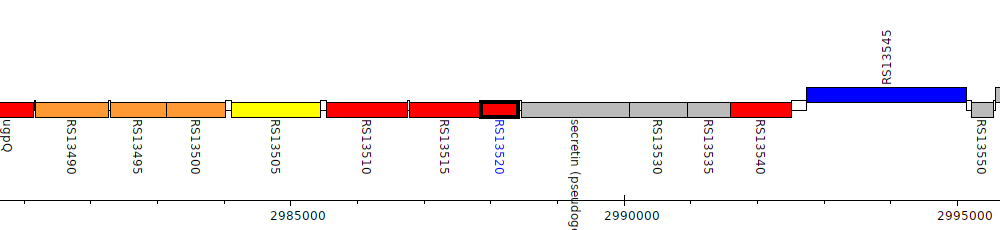
Burkholderia cenocepacia AU 1054, BCEN_RS13520
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | chorismate biosynthesis from 3-dehydroquinate | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00400 | Phenylalanine, tyrosine and tryptophan biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR01100 | Shikimate kinase family signature | 62 | 70 | 6.0E-23 | ||
| ProSitePatterns | PS01128 | Shikimate kinase signature. | IPR023000 | Shikimate kinase, conserved site | 63 | 88 | - |
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 9 | 171 | 1.8E-31 | |
| PRINTS | PR01100 | Shikimate kinase family signature | 80 | 89 | 6.0E-23 | ||
| PRINTS | PR01100 | Shikimate kinase family signature | 101 | 118 | 6.0E-23 | ||
| Pfam | PF01202 | Shikimate kinase | IPR031322 | Shikimate kinase/gluconokinase | 16 | 172 | 5.7E-52 |
| CDD | cd00464 | SK | IPR000623 | Shikimate kinase/Threonine synthase-like 1 | 9 | 161 | 2.87178E-74 |
| PRINTS | PR01100 | Shikimate kinase family signature | 34 | 47 | 6.0E-23 | ||
| Gene3D | G3DSA:3.40.50.300 | 2 | 174 | 3.9E-56 | |||
| Hamap | MF_00109 | Shikimate kinase [aroK]. | IPR000623 | Shikimate kinase/Threonine synthase-like 1 | 7 | 173 | 29.915 |
| PRINTS | PR01100 | Shikimate kinase family signature | 10 | 25 | 6.0E-23 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.