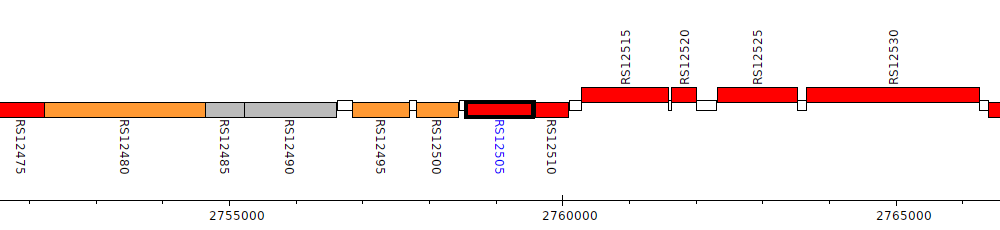
Burkholderia cenocepacia AU 1054, BCEN_RS12505
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009058 | biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:SSF53328
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0071951 | conversion of methionyl-tRNA to N-formyl-methionyl-tRNA |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00182
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF50486
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004479 | methionyl-tRNA formyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00182
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016742 | hydroxymethyl-, formyl- and related transferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF53328
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00970 | Aminoacyl-tRNA biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00670 | One carbon pool by folate | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd08704 | Met_tRNA_FMT_C | 221 | 309 | 2.98259E-30 | ||
| Pfam | PF02911 | Formyl transferase, C-terminal domain | IPR005793 | Formyl transferase, C-terminal | 216 | 316 | 4.4E-25 |
| SUPERFAMILY | SSF50486 | IPR011034 | Formyl transferase-like, C-terminal domain superfamily | 217 | 324 | 4.71E-29 | |
| Pfam | PF00551 | Formyl transferase | IPR002376 | Formyl transferase, N-terminal | 7 | 192 | 2.4E-41 |
| SUPERFAMILY | SSF53328 | IPR036477 | Formyl transferase, N-terminal domain superfamily | 4 | 216 | 4.71E-61 | |
| ProSitePatterns | PS00373 | Phosphoribosylglycinamide formyltransferase active site. | IPR001555 | Phosphoribosylglycinamide formyltransferase, active site | 144 | 167 | - |
| TIGRFAM | TIGR00460 | fmt: methionyl-tRNA formyltransferase | IPR005794 | Methionyl-tRNA formyltransferase | 5 | 317 | 2.4E-84 |
| Gene3D | G3DSA:3.10.25.10 | IPR037022 | Formyl transferase, C-terminal domain superfamily | 219 | 324 | 3.0E-27 | |
| CDD | cd08646 | FMT_core_Met-tRNA-FMT_N | IPR041711 | Methionyl-tRNA formyltransferase, N-terminal domain | 5 | 217 | 9.73426E-98 |
| Gene3D | G3DSA:3.40.50.170 | 1 | 218 | 8.7E-74 | |||
| Hamap | MF_00182 | Methionyl-tRNA formyltransferase [fmt]. | IPR005794 | Methionyl-tRNA formyltransferase | 5 | 319 | 34.865 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.