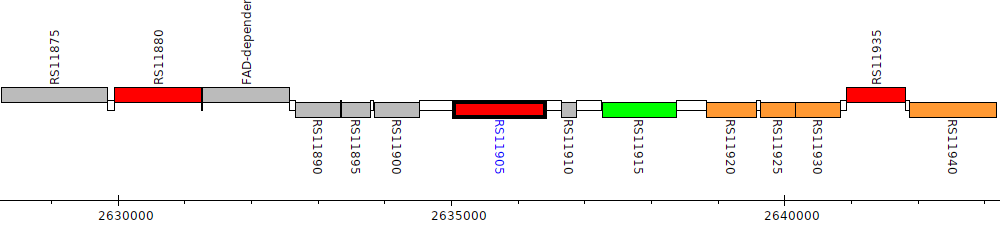
Burkholderia cenocepacia AU 1054, BCEN_RS11905
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016668 | oxidoreductase activity, acting on a sulfur group of donors, NAD(P) as acceptor |
Inferred from Sequence Model
Term mapped from: InterPro:PS00076
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF07992
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0050661 | NADP binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01424
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PF02852
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006749 | glutathione metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01424
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0050660 | flavin adenine dinucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF55424
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0009055 | electron transfer activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000350
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0045454 | cell redox homeostasis |
Inferred from Sequence Model
Term mapped from: InterPro:PF02852
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004362 | glutathione-disulfide reductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01424
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | glutathione-peroxide redox reactions | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00480 | Glutathione metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.50.50.60 | IPR036188 | FAD/NAD(P)-binding domain superfamily | 143 | 260 | 7.1E-113 | |
| Gene3D | G3DSA:3.50.50.60 | IPR036188 | FAD/NAD(P)-binding domain superfamily | 5 | 319 | 7.1E-113 | |
| PRINTS | PR00411 | Pyridine nucleotide disulphide reductase class-I signature | 168 | 193 | 2.2E-68 | ||
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | 282 | 304 | 2.8E-30 | ||
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | 131 | 149 | 2.8E-30 | ||
| PRINTS | PR00411 | Pyridine nucleotide disulphide reductase class-I signature | 134 | 143 | 2.2E-68 | ||
| Pfam | PF07992 | Pyridine nucleotide-disulphide oxidoreductase | IPR023753 | FAD/NAD(P)-binding domain | 5 | 317 | 8.5E-66 |
| PRINTS | PR00411 | Pyridine nucleotide disulphide reductase class-I signature | 254 | 268 | 2.2E-68 | ||
| PRINTS | PR00411 | Pyridine nucleotide disulphide reductase class-I signature | 297 | 304 | 2.2E-68 | ||
| PIRSF | PIRSF000350 | IPR001100 | Pyridine nucleotide-disulphide oxidoreductase, class I | 1 | 446 | 6.5E-93 | |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | 253 | 269 | 2.8E-30 | ||
| SUPERFAMILY | SSF55424 | IPR016156 | FAD/NAD-linked reductase, dimerisation domain superfamily | 335 | 449 | 1.89E-36 | |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | 7 | 26 | 2.8E-30 | ||
| SUPERFAMILY | SSF51905 | IPR036188 | FAD/NAD(P)-binding domain superfamily | 1 | 364 | 4.91E-57 | |
| PRINTS | PR00411 | Pyridine nucleotide disulphide reductase class-I signature | 333 | 354 | 2.2E-68 | ||
| ProSitePatterns | PS00076 | Pyridine nucleotide-disulphide oxidoreductases class-I active site. | IPR012999 | Pyridine nucleotide-disulphide oxidoreductase, class I, active site | 39 | 49 | - |
| Gene3D | G3DSA:3.30.390.30 | IPR016156 | FAD/NAD-linked reductase, dimerisation domain superfamily | 336 | 451 | 2.2E-45 | |
| PRINTS | PR00411 | Pyridine nucleotide disulphide reductase class-I signature | 38 | 53 | 2.2E-68 | ||
| PRINTS | PR00411 | Pyridine nucleotide disulphide reductase class-I signature | 398 | 413 | 2.2E-68 | ||
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | 168 | 186 | 2.8E-30 | ||
| TIGRFAM | TIGR01424 | gluta_reduc_2: glutathione-disulfide reductase | IPR006324 | Glutathione-disulphide reductase | 3 | 446 | 1.4E-188 |
| PRINTS | PR00411 | Pyridine nucleotide disulphide reductase class-I signature | 420 | 440 | 2.2E-68 | ||
| Pfam | PF02852 | Pyridine nucleotide-disulphide oxidoreductase, dimerisation domain | IPR004099 | Pyridine nucleotide-disulphide oxidoreductase, dimerisation domain | 337 | 445 | 1.4E-35 |
| PRINTS | PR00411 | Pyridine nucleotide disulphide reductase class-I signature | 6 | 28 | 2.2E-68 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.