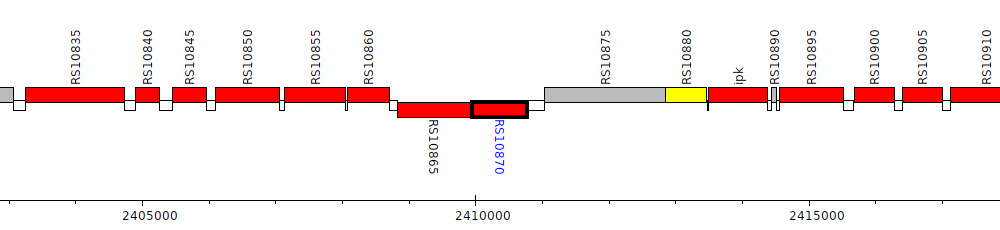
Burkholderia cenocepacia AU 1054, BCEN_RS10870
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006284 | base-excision repair |
Inferred from Sequence Model
Term mapped from: InterPro:SM00898
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003676 | nucleic acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF46946
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006281 | DNA repair |
Inferred from Sequence Model
Term mapped from: InterPro:PS01242
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016799 | hydrolase activity, hydrolyzing N-glycosyl compounds |
Inferred from Sequence Model
Term mapped from: InterPro:SM00898
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003684 | damaged DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00898
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008534 | oxidized purine nucleobase lesion DNA N-glycosylase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00103
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003906 | DNA-(apurinic or apyrimidinic site) endonuclease activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00898
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008270 | zinc ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00898
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS01242
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006289 | nucleotide-excision repair |
Inferred from Sequence Model
Term mapped from: InterPro:PF06831
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00898 | IPR012319 | Formamidopyrimidine-DNA glycosylase, catalytic domain | 2 | 117 | 1.1E-46 | |
| Pfam | PF01149 | Formamidopyrimidine-DNA glycosylase N-terminal domain | IPR012319 | Formamidopyrimidine-DNA glycosylase, catalytic domain | 1 | 116 | 6.2E-31 |
| Gene3D | G3DSA:1.10.8.50 | 145 | 275 | 1.3E-43 | |||
| CDD | cd08966 | EcFpg-like_N | 2 | 120 | 3.59311E-45 | ||
| Gene3D | G3DSA:3.20.190.10 | IPR035937 | MutM-like, N-terminal | 2 | 129 | 4.4E-35 | |
| TIGRFAM | TIGR00577 | fpg: DNA-formamidopyrimidine glycosylase | IPR020629 | Formamidopyrimidine-DNA glycosylase | 1 | 274 | 2.8E-84 |
| Pfam | PF06827 | Zinc finger found in FPG and IleRS | IPR010663 | Zinc finger, FPG/IleRS-type | 247 | 274 | 4.2E-11 |
| SUPERFAMILY | SSF46946 | IPR010979 | Ribosomal protein S13-like, H2TH | 136 | 227 | 1.3E-25 | |
| ProSiteProfiles | PS51066 | Zinc finger FPG-type profile. | IPR000214 | Zinc finger, DNA glycosylase/AP lyase-type | 241 | 275 | 13.976 |
| ProSitePatterns | PS01242 | Zinc finger FPG-type signature. | IPR015887 | DNA glycosylase/AP lyase, zinc finger domain, DNA-binding site | 250 | 274 | - |
| Hamap | MF_00103 | Formamidopyrimidine-DNA glycosylase [mutM]. | IPR020629 | Formamidopyrimidine-DNA glycosylase | 1 | 275 | 42.426 |
| SMART | SM01232 | IPR015886 | DNA glycosylase/AP lyase, H2TH DNA-binding | 135 | 227 | 4.0E-43 | |
| Pfam | PF06831 | Formamidopyrimidine-DNA glycosylase H2TH domain | IPR015886 | DNA glycosylase/AP lyase, H2TH DNA-binding | 135 | 225 | 2.7E-22 |
| SUPERFAMILY | SSF57716 | 223 | 274 | 1.14E-17 | |||
| ProSiteProfiles | PS51068 | Formamidopyrimidine-DNA glycosylase catalytic domain profile. | IPR012319 | Formamidopyrimidine-DNA glycosylase, catalytic domain | 2 | 114 | 31.636 |
| SUPERFAMILY | SSF81624 | IPR035937 | MutM-like, N-terminal | 2 | 144 | 8.63E-37 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.