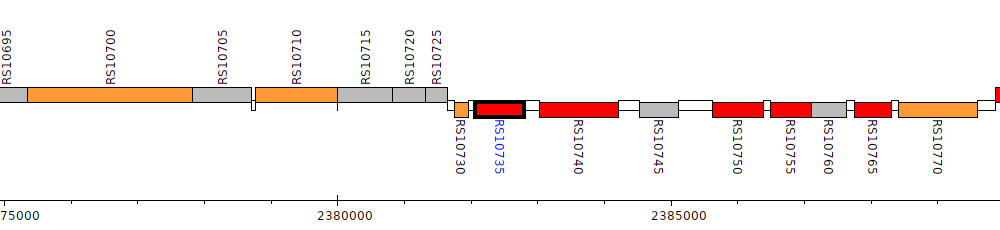
Burkholderia cenocepacia AU 1054, BCEN_RS10735
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS00061
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0018454 | acetoacetyl-CoA reductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01829
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0042619 | poly-hydroxybutyrate biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01829
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005737 | cytoplasm |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01829
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01829
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | acetyl-CoA fermentation to butanoate II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00630 | Glyoxylate and dicarboxylate metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | (R)- and (S)-3-hydroxybutanoate biosynthesis (engineered) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | ethylmalonyl-CoA pathway | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00650 | Butanoate metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | 4-oxopentanoate degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00822 | This enzymatic domain is part of bacterial polyketide synthases and catalyses the first step in the reductive modification of the beta-carbonyl centres in the growing polyketide chain. It uses NADPH to reduce the keto group to a hydroxy group. | 4 | 184 | 5.7E-5 | ||
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 174 | 191 | 5.1E-48 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 80 | 91 | 5.1E-48 |
| PRINTS | PR00080 | Short-chain dehydrogenase/reductase (SDR) superfamily signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 133 | 141 | 2.7E-16 |
| ProSitePatterns | PS00061 | Short-chain dehydrogenases/reductases family signature. | IPR020904 | Short-chain dehydrogenase/reductase, conserved site | 140 | 168 | - |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 127 | 143 | 5.1E-48 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 208 | 228 | 5.1E-48 |
| TIGRFAM | TIGR01829 | AcAcCoA_reduct: acetoacetyl-CoA reductase | IPR011283 | Acetoacetyl-CoA reductase | 4 | 246 | 4.1E-110 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 5 | 22 | 5.1E-48 |
| SUPERFAMILY | SSF51735 | IPR036291 | NAD(P)-binding domain superfamily | 3 | 246 | 5.88E-79 | |
| PRINTS | PR00080 | Short-chain dehydrogenase/reductase (SDR) superfamily signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 153 | 172 | 2.7E-16 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 153 | 172 | 5.1E-48 |
| CDD | cd05333 | BKR_SDR_c | 4 | 245 | 1.33491E-122 | ||
| Pfam | PF13561 | Enoyl-(Acyl carrier protein) reductase | 13 | 245 | 1.0E-60 | ||
| PRINTS | PR00080 | Short-chain dehydrogenase/reductase (SDR) superfamily signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 80 | 91 | 2.7E-16 |
| Gene3D | G3DSA:3.40.50.720 | 1 | 247 | 7.3E-84 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.