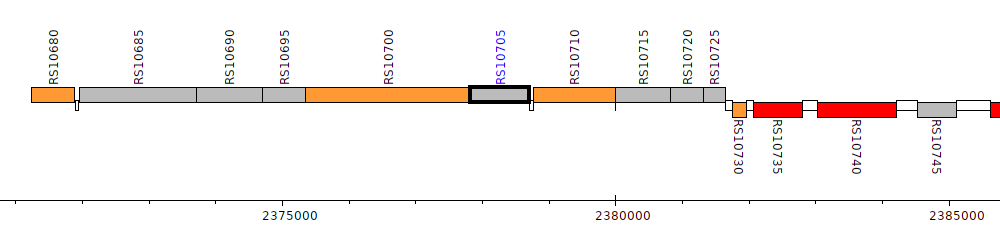
Burkholderia cenocepacia AU 1054, BCEN_RS10705
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016810 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds |
Inferred from Sequence Model
Term mapped from: InterPro:PF01522
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF88713
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0005975 | carbohydrate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF01522
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| ProSiteProfiles | PS51677 | NodB homology domain profile. | IPR002509 | NodB homology domain | 84 | 294 | 25.23 |
| CDD | cd10917 | CE4_NodB_like_6s_7s | 84 | 261 | 3.43855E-54 | ||
| SUPERFAMILY | SSF88713 | IPR011330 | Glycoside hydrolase/deacetylase, beta/alpha-barrel | 54 | 274 | 3.11E-49 | |
| Pfam | PF01522 | Polysaccharide deacetylase | IPR002509 | NodB homology domain | 83 | 201 | 3.2E-26 |
| Gene3D | G3DSA:3.20.20.370 | 81 | 279 | 1.4E-41 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.