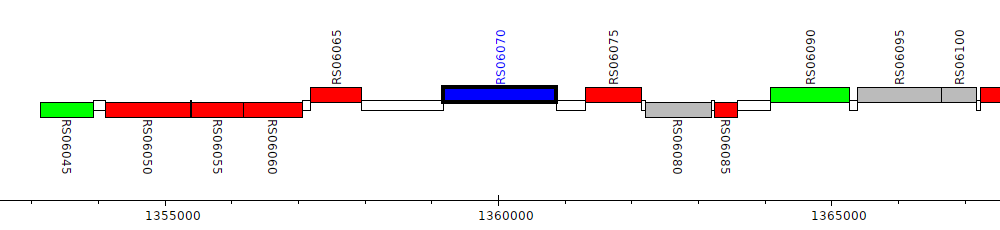
Burkholderia cenocepacia AU 1054, BCEN_RS06070
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004222 | metalloendopeptidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00730
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006508 | proteolysis |
Inferred from Sequence Model
Term mapped from: InterPro:PR00730
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:1.10.390.10 | IPR027268 | Peptidase M4/M1, CTD superfamily | 389 | 564 | 7.1E-47 | |
| PRINTS | PR00730 | Thermolysin metalloprotease (M4) family signature | IPR023612 | Peptidase M4 | 396 | 407 | 2.6E-15 |
| PRINTS | PR00730 | Thermolysin metalloprotease (M4) family signature | IPR023612 | Peptidase M4 | 338 | 358 | 2.6E-15 |
| PRINTS | PR00730 | Thermolysin metalloprotease (M4) family signature | IPR023612 | Peptidase M4 | 375 | 391 | 2.6E-15 |
| Pfam | PF01447 | Thermolysin metallopeptidase, catalytic domain | IPR013856 | Peptidase M4 domain | 222 | 384 | 5.8E-35 |
| Pfam | PF02868 | Thermolysin metallopeptidase, alpha-helical domain | IPR001570 | Peptidase M4, C-terminal | 387 | 564 | 2.4E-49 |
| Pfam | PF03413 | Peptidase propeptide and YPEB domain | IPR025711 | PepSY domain | 148 | 211 | 6.9E-5 |
| Pfam | PF07504 | Fungalysin/Thermolysin Propeptide Motif | IPR011096 | FTP domain | 70 | 116 | 1.4E-14 |
| Gene3D | G3DSA:3.10.450.490 | 33 | 125 | 3.2E-5 | |||
| Gene3D | G3DSA:3.10.170.10 | 221 | 388 | 2.1E-35 | |||
| CDD | cd09597 | M4_TLP | 256 | 564 | 3.45578E-100 | ||
| Coils | Coil | 564 | 565 | - | |||
| PRINTS | PR00730 | Thermolysin metalloprotease (M4) family signature | IPR023612 | Peptidase M4 | 464 | 480 | 2.6E-15 |
| SUPERFAMILY | SSF55486 | 219 | 564 | 1.1E-62 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.