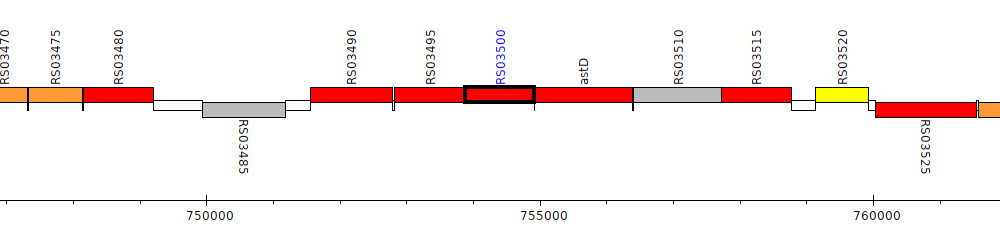
Burkholderia cenocepacia AU 1054, BCEN_RS03500
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006527 | arginine catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03243
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008791 | arginine N-succinyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03243
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00330 | Arginine and proline metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF04958 | Arginine N-succinyltransferase beta subunit | IPR007041 | Arginine N-succinyltransferase AstA/AruG | 1 | 337 | 1.8E-135 |
| TIGRFAM | TIGR03244 | arg_catab_AstA: arginine N-succinyltransferase | IPR017650 | Arginine N-succinyltransferase | 3 | 338 | 2.7E-151 |
| SUPERFAMILY | SSF55729 | IPR016181 | Acyl-CoA N-acyltransferase | 1 | 338 | 2.08E-125 | |
| Gene3D | G3DSA:2.40.40.20 | 273 | 339 | 2.1E-5 | |||
| Gene3D | G3DSA:3.40.630.30 | 1 | 272 | 2.4E-116 | |||
| TIGRFAM | TIGR03243 | arg_catab_AOST: arginine and ornithine succinyltransferase subunits | IPR007041 | Arginine N-succinyltransferase AstA/AruG | 3 | 338 | 2.2E-141 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.