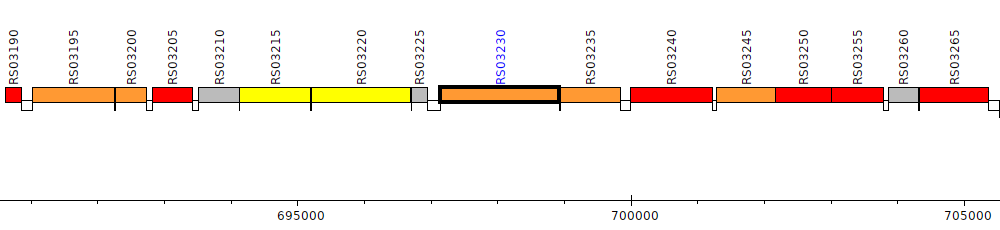
Burkholderia cenocepacia AU 1054, BCEN_RS03230
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0005525 | GTP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00009
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003924 | GTPase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00009
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF03144 | Elongation factor Tu domain 2 | IPR004161 | Translation elongation factor EFTu-like, domain 2 | 205 | 275 | 5.0E-10 |
| ProSiteProfiles | PS51722 | Translational (tr)-type guanine nucleotide-binding (G) domain profile. | IPR000795 | Transcription factor, GTP-binding domain | 2 | 184 | 58.79 |
| Pfam | PF00679 | Elongation factor G C-terminus | IPR000640 | Elongation factor EFG, domain V-like | 399 | 483 | 3.4E-23 |
| CDD | cd03699 | EF4_II | 191 | 276 | 7.95991E-42 | ||
| CDD | cd01890 | LepA | 5 | 183 | 3.53409E-124 | ||
| Gene3D | G3DSA:3.30.70.2570 | IPR038363 | LepA, C-terminal domain superfamily | 479 | 553 | 3.2E-32 | |
| SMART | SM00838 | IPR000640 | Elongation factor EFG, domain V-like | 401 | 486 | 0.0097 | |
| PRINTS | PR00315 | GTP-binding elongation factor signature | IPR000795 | Transcription factor, GTP-binding domain | 126 | 135 | 2.3E-14 |
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 2 | 205 | 2.39E-55 | |
| SUPERFAMILY | SSF54980 | IPR035647 | EF-G domain III/V-like | 399 | 493 | 5.76E-26 | |
| TIGRFAM | TIGR00231 | small_GTP: small GTP-binding protein domain | IPR005225 | Small GTP-binding protein domain | 5 | 172 | 4.5E-17 |
| PRINTS | PR00315 | GTP-binding elongation factor signature | IPR000795 | Transcription factor, GTP-binding domain | 74 | 84 | 2.3E-14 |
| Pfam | PF00009 | Elongation factor Tu GTP binding domain | IPR000795 | Transcription factor, GTP-binding domain | 3 | 181 | 4.6E-52 |
| CDD | cd03709 | lepA_C | IPR035654 | Elongation factor 4, domain IV | 401 | 479 | 1.29254E-41 |
| CDD | cd16260 | EF4_III | 293 | 368 | 5.28433E-49 | ||
| Gene3D | G3DSA:3.40.50.300 | 1 | 185 | 2.4E-69 | |||
| Pfam | PF06421 | GTP-binding protein LepA C-terminus | IPR013842 | GTP-binding protein LepA, C-terminal | 487 | 593 | 8.4E-50 |
| Gene3D | G3DSA:2.40.30.10 | 186 | 285 | 4.5E-35 | |||
| SUPERFAMILY | SSF50447 | IPR009000 | Translation protein, beta-barrel domain superfamily | 165 | 288 | 5.74E-29 | |
| PRINTS | PR00315 | GTP-binding elongation factor signature | IPR000795 | Transcription factor, GTP-binding domain | 49 | 57 | 2.3E-14 |
| TIGRFAM | TIGR01393 | lepA: elongation factor 4 | IPR006297 | Elongation factor 4 | 3 | 595 | 2.4E-300 |
| PRINTS | PR00315 | GTP-binding elongation factor signature | IPR000795 | Transcription factor, GTP-binding domain | 6 | 19 | 2.3E-14 |
| PRINTS | PR00315 | GTP-binding elongation factor signature | IPR000795 | Transcription factor, GTP-binding domain | 90 | 101 | 2.3E-14 |
| ProSitePatterns | PS00301 | Translational (tr)-type guanine nucleotide-binding (G) domain signature. | IPR031157 | Tr-type G domain, conserved site | 42 | 57 | - |
| Gene3D | G3DSA:3.30.70.870 | 286 | 367 | 2.5E-36 | |||
| Hamap | MF_00071 | Elongation factor 4 [lepA]. | IPR006297 | Elongation factor 4 | 1 | 597 | 306.886 |
| Gene3D | G3DSA:3.30.70.3380 | 368 | 478 | 8.2E-43 | |||
| SUPERFAMILY | SSF54980 | IPR035647 | EF-G domain III/V-like | 292 | 370 | 5.76E-18 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.