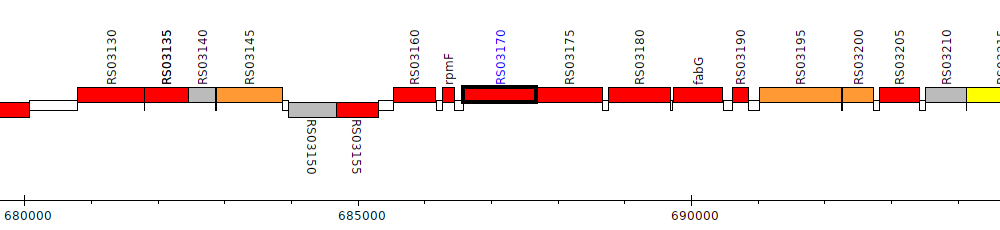
Burkholderia cenocepacia AU 1054, BCEN_RS03170
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016747 | transferase activity, transferring acyl groups other than amino-acyl groups |
Inferred from Sequence Model
Term mapped from: InterPro:PF02504
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006633 | fatty acid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF02504
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF02504
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF53659 | 4 | 332 | 2.75E-105 | |||
| PIRSF | PIRSF002465 | IPR012281 | Phospholipid biosynthesis protein, PlsX-like | 1 | 336 | 1.0E-130 | |
| TIGRFAM | TIGR00182 | plsX: fatty acid/phospholipid synthesis protein PlsX | IPR012281 | Phospholipid biosynthesis protein, PlsX-like | 4 | 331 | 7.2E-109 |
| Pfam | PF02504 | Fatty acid synthesis protein | IPR003664 | Fatty acid synthesis PlsX protein | 3 | 325 | 6.3E-111 |
| Gene3D | G3DSA:3.40.718.10 | 1 | 345 | 1.8E-127 | |||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 338 | 368 | - | ||
| Hamap | MF_00019 | Phosphate acyltransferase [plsX]. | IPR012281 | Phospholipid biosynthesis protein, PlsX-like | 1 | 338 | 155.985 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.