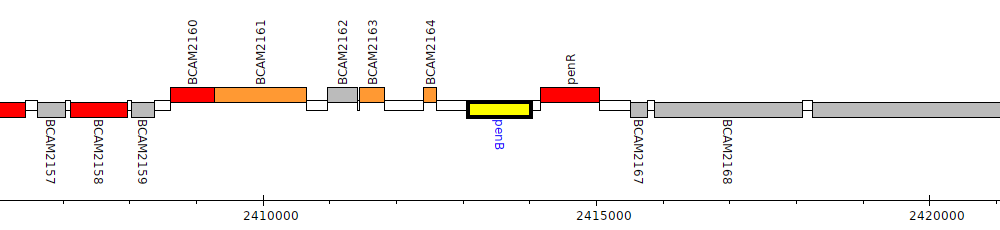
Burkholderia cenocepacia J2315, BCAM2165 (penB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0008800 | beta-lactamase activity | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0046677 | response to antibiotic |
Inferred from Sequence Model
Term mapped from: InterPro:PR00118
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0030655 | beta-lactam antibiotic catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PR00118
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008800 | beta-lactamase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00118
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00311 | Penicillin and cephalosporin biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00311 | Penicillin and cephalosporin biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01501 | beta-Lactam resistance | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00118 | Beta-lactamase class A signature | IPR000871 | Beta-lactamase, class-A | 244 | 259 | 3.4E-75 |
| PRINTS | PR00118 | Beta-lactamase class A signature | IPR000871 | Beta-lactamase, class-A | 191 | 216 | 3.4E-75 |
| ProSitePatterns | PS00146 | Beta-lactamase class-A active site. | IPR023650 | Beta-lactamase, class-A active site | 89 | 104 | - |
| PRINTS | PR00118 | Beta-lactamase class A signature | IPR000871 | Beta-lactamase, class-A | 165 | 189 | 3.4E-75 |
| PRINTS | PR00118 | Beta-lactamase class A signature | IPR000871 | Beta-lactamase, class-A | 129 | 154 | 3.4E-75 |
| PRINTS | PR00118 | Beta-lactamase class A signature | IPR000871 | Beta-lactamase, class-A | 87 | 104 | 3.4E-75 |
| PRINTS | PR00118 | Beta-lactamase class A signature | IPR000871 | Beta-lactamase, class-A | 56 | 80 | 3.4E-75 |
| SUPERFAMILY | SSF56601 | IPR012338 | Beta-lactamase/transpeptidase-like | 56 | 310 | 1.67E-81 | |
| Pfam | PF13354 | Beta-lactamase enzyme family | 77 | 284 | 1.9E-42 | ||
| PRINTS | PR00118 | Beta-lactamase class A signature | IPR000871 | Beta-lactamase, class-A | 227 | 242 | 3.4E-75 |
| ProSiteProfiles | PS51318 | Twin arginine translocation (Tat) signal profile. | IPR006311 | Twin-arginine translocation pathway, signal sequence | 1 | 33 | 8.12 |
| Gene3D | G3DSA:3.40.710.10 | 49 | 311 | 2.9E-85 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.