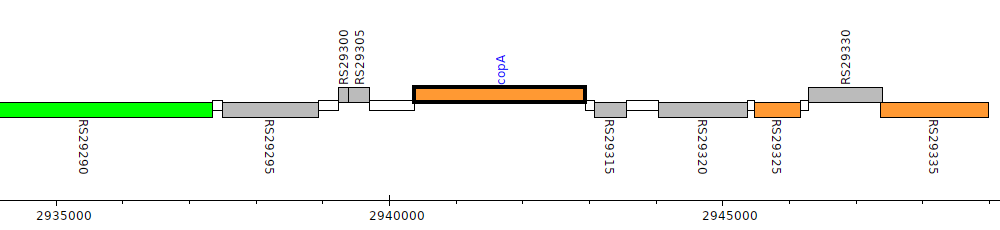
Burkholderia cenocepacia H111, I35_RS29310 (copA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0030001 | metal ion transport |
Inferred from Sequence Model
Term mapped from: InterPro:PF00403
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0019829 | ATPase-coupled cation transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01525
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016021 | integral component of membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01494
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000166 | nucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.1110.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005507 | copper ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00003
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006812 | cation transport |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01525
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0046872 | metal ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00403
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | 693 | 703 | 3.4E-24 | ||
| Gene3D | G3DSA:3.40.1110.10 | IPR023299 | P-type ATPase, cytoplasmic domain N | 549 | 678 | 1.1E-81 | |
| SUPERFAMILY | SSF81665 | IPR023298 | P-type ATPase, transmembrane domain superfamily | 300 | 824 | 5.36E-15 | |
| Gene3D | G3DSA:3.30.70.100 | 112 | 177 | 1.4E-20 | |||
| CDD | cd00371 | HMA | IPR006121 | Heavy metal-associated domain, HMA | 32 | 92 | 8.23004E-22 |
| ProSitePatterns | PS01047 | Heavy-metal-associated domain. | IPR017969 | Heavy-metal-associated, conserved site | 116 | 145 | - |
| ProSitePatterns | PS01047 | Heavy-metal-associated domain. | IPR017969 | Heavy-metal-associated, conserved site | 35 | 64 | - |
| PRINTS | PR00942 | Copper-transporting ATPase 1 signature | 30 | 55 | 6.9E-9 | ||
| ProSiteProfiles | PS50846 | Heavy-metal-associated domain profile. | IPR006121 | Heavy metal-associated domain, HMA | 111 | 176 | 19.685 |
| CDD | cd00371 | HMA | IPR006121 | Heavy metal-associated domain, HMA | 113 | 175 | 1.42751E-14 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | 538 | 552 | 3.4E-24 | ||
| SUPERFAMILY | SSF81653 | IPR008250 | P-type ATPase, A domain superfamily | 341 | 438 | 5.49E-26 | |
| Pfam | PF00122 | E1-E2 ATPase | 339 | 518 | 2.4E-56 | ||
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | 388 | 402 | 3.4E-24 | ||
| TIGRFAM | TIGR00003 | TIGR00003: copper ion binding protein | IPR006122 | Heavy metal-associated domain, copper ion-binding | 31 | 92 | 4.4E-13 |
| TIGRFAM | TIGR01494 | ATPase_P-type: HAD ATPase, P-type, family IC | IPR001757 | P-type ATPase | 310 | 558 | 1.3E-45 |
| TIGRFAM | TIGR01525 | ATPase-IB_hvy: heavy metal translocating P-type ATPase | IPR027256 | P-type ATPase, subfamily IB | 274 | 849 | 3.4E-186 |
| SFLD | SFLDG00002 | C1.7: P-type atpase like | 520 | 798 | 1.261E-44 | ||
| Pfam | PF00403 | Heavy-metal-associated domain | IPR006121 | Heavy metal-associated domain, HMA | 33 | 91 | 3.1E-17 |
| SUPERFAMILY | SSF55008 | IPR036163 | Heavy metal-associated domain superfamily | 110 | 176 | 7.98E-19 | |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 1 | 20 | - | ||
| Pfam | PF00403 | Heavy-metal-associated domain | IPR006121 | Heavy metal-associated domain, HMA | 114 | 173 | 2.7E-14 |
| SFLD | SFLDF00027 | p-type atpase | 520 | 798 | 1.261E-44 | ||
| Pfam | PF00702 | haloacid dehalogenase-like hydrolase | 535 | 761 | 1.2E-40 | ||
| Gene3D | G3DSA:2.70.150.20 | 326 | 442 | 1.9E-37 | |||
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | 770 | 782 | 3.4E-24 | ||
| SUPERFAMILY | SSF56784 | IPR036412 | HAD-like superfamily | 536 | 847 | 1.46E-53 | |
| CDD | cd02094 | P-type_ATPase_Cu-like | 214 | 851 | 0.0 | ||
| ProSiteProfiles | PS50846 | Heavy-metal-associated domain profile. | IPR006121 | Heavy metal-associated domain, HMA | 30 | 95 | 22.756 |
| PRINTS | PR00942 | Copper-transporting ATPase 1 signature | 257 | 282 | 6.9E-9 | ||
| PRINTS | PR00942 | Copper-transporting ATPase 1 signature | 134 | 155 | 6.9E-9 | ||
| Gene3D | G3DSA:3.40.50.1000 | IPR023214 | HAD superfamily | 519 | 799 | 1.1E-81 | |
| Gene3D | G3DSA:3.30.70.100 | 29 | 96 | 4.2E-23 | |||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 183 | 203 | - | ||
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | 747 | 766 | 3.4E-24 | ||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 1 | 25 | - | ||
| ProSitePatterns | PS00154 | E1-E2 ATPases phosphorylation site. | IPR018303 | P-type ATPase, phosphorylation site | 540 | 546 | - |
| TIGRFAM | TIGR01494 | ATPase_P-type: HAD ATPase, P-type, family IC | IPR001757 | P-type ATPase | 667 | 829 | 6.0E-37 |
| SUPERFAMILY | SSF55008 | IPR036163 | Heavy metal-associated domain superfamily | 29 | 95 | 2.75E-21 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.