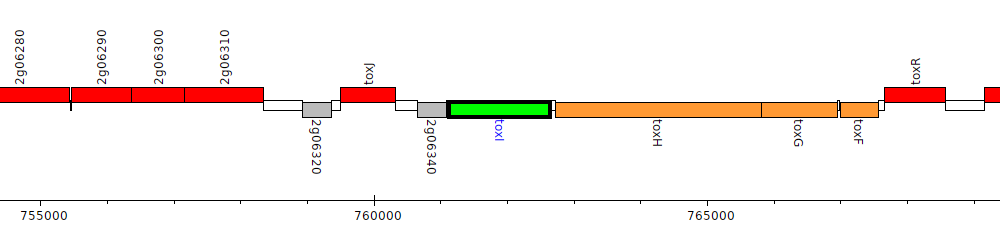
Burkholderia glumae BGR1, bglu_2g06350 (toxI)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01845
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0022857 | transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01845
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0015562 | efflux transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF02321
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055085 | transmembrane transport |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01845
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bgl01501 | beta-Lactam resistance | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| TIGRFAM | TIGR01845 | outer_NodT: efflux transporter, outer membrane factor (OMF) lipoprotein, NodT family | IPR010131 | RND efflux system, outer membrane lipoprotein, NodT | 5 | 467 | 4.9E-122 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 472 | 506 | - | ||
| Gene3D | G3DSA:2.20.200.10 | 104 | 356 | 1.7E-141 | |||
| Gene3D | G3DSA:1.20.1600.10 | 58 | 467 | 1.7E-141 | |||
| Pfam | PF02321 | Outer membrane efflux protein | IPR003423 | Outer membrane efflux protein | 286 | 465 | 3.0E-26 |
| SUPERFAMILY | SSF56954 | 3 | 469 | 3.92E-116 | |||
| Pfam | PF02321 | Outer membrane efflux protein | IPR003423 | Outer membrane efflux protein | 67 | 256 | 9.2E-30 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.