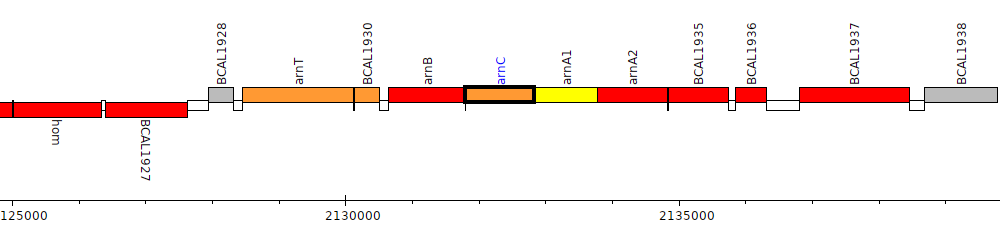
Burkholderia cenocepacia J2315, BCAL1932 (arnC)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009245 | lipid A biosynthetic process |
Inferred from Sequence or Structural Similarity
Term mapped from: UniprotKB:O52324
|
ECO:0000044 sequence similarity evidence |
17337576 | Reviewed by curator |
| Biological Process | GO:1901760 | beta-L-Ara4N-lipid A biosynthetic process | Inferred from Genetic Interaction | ECO:0006095 |
22742453 | Reviewed by curator |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj00520 | Amino sugar and nucleotide sugar metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF53448 | IPR029044 | Nucleotide-diphospho-sugar transferases | 15 | 228 | 1.88E-34 | |
| CDD | cd04187 | DPM1_like_bac | 25 | 202 | 7.1633E-75 | ||
| Pfam | PF00535 | Glycosyl transferase family 2 | IPR001173 | Glycosyltransferase 2-like | 24 | 184 | 8.9E-28 |
| Gene3D | G3DSA:3.90.550.10 | IPR029044 | Nucleotide-diphospho-sugar transferases | 8 | 225 | 2.7E-36 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.