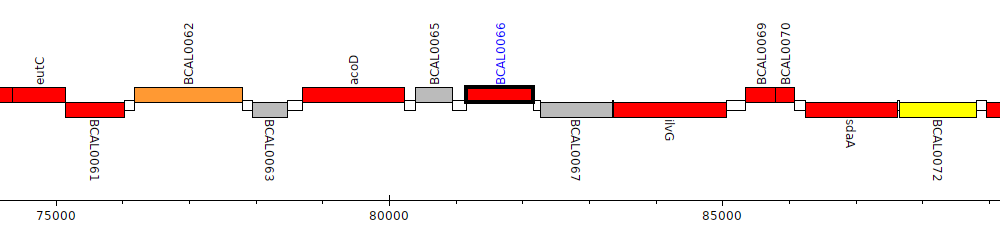
Burkholderia cenocepacia J2315, BCAL0066
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF46689
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003700 | DNA-binding transcription factor activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00342
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0043565 | sequence-specific DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00342
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:SM00342
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:1.10.10.60 | 235 | 338 | 6.7E-34 | |||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 1 | 21 | - | ||
| Pfam | PF12833 | Helix-turn-helix domain | IPR018060 | DNA binding HTH domain, AraC-type | 256 | 335 | 5.8E-28 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 1 | 22 | - | ||
| SUPERFAMILY | SSF46689 | IPR009057 | Homeobox-like domain superfamily | 286 | 335 | 4.84E-5 | |
| SMART | SM00342 | IPR018060 | DNA binding HTH domain, AraC-type | 250 | 334 | 4.1E-23 | |
| SUPERFAMILY | SSF46689 | IPR009057 | Homeobox-like domain superfamily | 234 | 284 | 6.48E-8 | |
| ProSiteProfiles | PS01124 | Bacterial regulatory proteins, araC family DNA-binding domain profile. | IPR018060 | DNA binding HTH domain, AraC-type | 237 | 336 | 24.145 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.