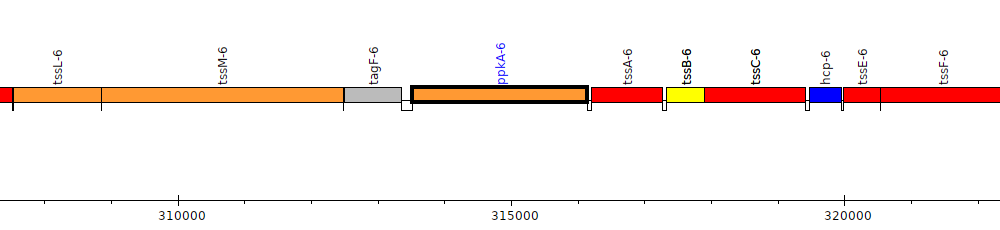
Burkholderia thailandensis E264 ATCC 700388, BTH_II0256 (ppkA-6)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0033103 | protein secretion by the type VI secretion system | Traceable Author Statement | ECO:0000304 traceable author statement used in manual assertion |
20865170 | Reviewed by curator |
| Biological Process | GO:0033103 | protein secretion by the type VI secretion system | Inferred from Genomic Context | ECO:0000177 genomic context evidence |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS50011
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006468 | protein phosphorylation |
Inferred from Sequence Model
Term mapped from: InterPro:PS50011
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004713 | protein tyrosine kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00219
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004672 | protein kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS50011
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| MobiDBLite | mobidb-lite | consensus disorder prediction | 467 | 482 | - | ||
| SMART | SM00219 | IPR020635 | Tyrosine-protein kinase, catalytic domain | 36 | 306 | 1.5E-8 | |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 447 | 462 | - | ||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 518 | 538 | - | ||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 330 | 362 | - | ||
| Gene3D | G3DSA:1.10.510.10 | 137 | 310 | 9.1E-50 | |||
| CDD | cd14014 | STKc_PknB_like | 37 | 310 | 7.33763E-61 | ||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 776 | 809 | - | ||
| Gene3D | G3DSA:3.30.200.20 | 5 | 136 | 9.1E-50 | |||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 501 | 515 | - | ||
| SUPERFAMILY | SSF56112 | IPR011009 | Protein kinase-like domain superfamily | 32 | 359 | 3.99E-53 | |
| ProSitePatterns | PS00109 | Tyrosine protein kinases specific active-site signature. | IPR008266 | Tyrosine-protein kinase, active site | 173 | 185 | - |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 323 | 569 | - | ||
| ProSitePatterns | PS00107 | Protein kinases ATP-binding region signature. | IPR017441 | Protein kinase, ATP binding site | 42 | 65 | - |
| Pfam | PF00069 | Protein kinase domain | IPR000719 | Protein kinase domain | 38 | 256 | 1.6E-28 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 667 | 713 | - | ||
| ProSiteProfiles | PS50011 | Protein kinase domain profile. | IPR000719 | Protein kinase domain | 36 | 312 | 29.333 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.