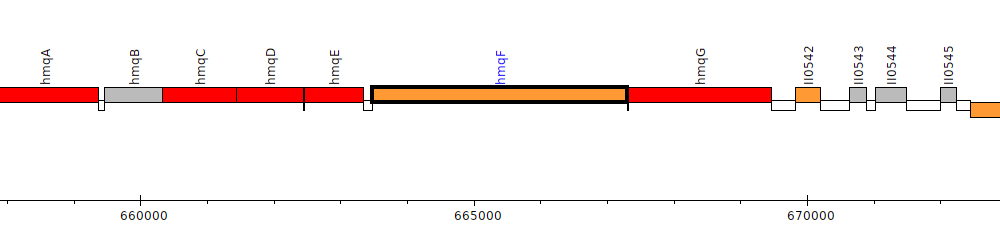
Burkholderia pseudomallei 1026b, BP1026B_II0540 (hmqF)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0050660 | flavin adenine dinucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF02771
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0031177 | phosphopantetheine binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00823
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00501
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:SSF56645
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016627 | oxidoreductase activity, acting on the CH-CH group of donors |
Inferred from Sequence Model
Term mapped from: InterPro:SSF56645
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0008610 | lipid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:cd05931
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0031177 | phosphopantetheine binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00823
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:SSF56645
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016627 | oxidoreductase activity, acting on the CH-CH group of donors |
Inferred from Sequence Model
Term mapped from: InterPro:SSF56645
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0008610 | lipid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:cd05931
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00501
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0050660 | flavin adenine dinucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF02771
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF47203 | IPR036250 | Acyl-CoA dehydrogenase-like, C-terminal | 844 | 997 | 3.06E-33 | |
| CDD | cd05931 | FAAL | IPR040097 | Fatty acyl-AMP ligase | 17 | 574 | 0.0 |
| Pfam | PF00550 | Phosphopantetheine attachment site | IPR009081 | Phosphopantetheine binding ACP domain | 1201 | 1252 | 4.4E-9 |
| Gene3D | G3DSA:1.10.540.10 | IPR037069 | Acyl-CoA dehydrogenase/oxidase, N-terminal domain superfamily | 629 | 738 | 2.1E-83 | |
| Gene3D | G3DSA:1.10.1200.10 | IPR036736 | ACP-like superfamily | 1171 | 1258 | 7.7E-16 | |
| SUPERFAMILY | SSF47336 | IPR036736 | ACP-like superfamily | 1179 | 1264 | 1.16E-13 | |
| Coils | Coil | 901 | 921 | - | |||
| Gene3D | G3DSA:3.40.50.12780 | IPR042099 | AMP-dependent synthetase-like superfamily | 2 | 465 | 1.2E-145 | |
| Pfam | PF02771 | Acyl-CoA dehydrogenase, N-terminal domain | IPR013786 | Acyl-CoA dehydrogenase/oxidase, N-terminal | 630 | 735 | 1.5E-14 |
| ProSitePatterns | PS00455 | Putative AMP-binding domain signature. | IPR020845 | AMP-binding, conserved site | 170 | 181 | - |
| ProSiteProfiles | PS50075 | Carrier protein (CP) domain profile. | IPR009081 | Phosphopantetheine binding ACP domain | 1178 | 1256 | 12.665 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 600 | 614 | - | ||
| Gene3D | G3DSA:3.30.300.30 | 466 | 595 | 7.5E-40 | |||
| SMART | SM01294 | 1186 | 1252 | 3.1E-6 | |||
| Pfam | PF02770 | Acyl-CoA dehydrogenase, middle domain | IPR006091 | Acyl-CoA oxidase/dehydrogenase, central domain | 740 | 831 | 3.0E-17 |
| Pfam | PF00501 | AMP-binding enzyme | IPR000873 | AMP-dependent synthetase/ligase | 16 | 471 | 7.1E-84 |
| Coils | Coil | 1277 | 1277 | - | |||
| Gene3D | G3DSA:1.20.140.10 | 845 | 986 | 2.1E-83 | |||
| Pfam | PF00441 | Acyl-CoA dehydrogenase, C-terminal domain | IPR009075 | Acyl-CoA dehydrogenase/oxidase C-terminal | 845 | 994 | 1.1E-31 |
| SMART | SM00823 | Phosphopantetheine attachment site | IPR020806 | Polyketide synthase, phosphopantetheine-binding domain | 1185 | 1256 | 7.4E-12 |
| SUPERFAMILY | SSF56645 | IPR009100 | Acyl-CoA dehydrogenase/oxidase, N-terminal and middle domain superfamily | 628 | 855 | 1.31E-45 | |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 1149 | 1178 | - | ||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 599 | 624 | - | ||
| ProSitePatterns | PS00012 | Phosphopantetheine attachment site. | IPR006162 | Phosphopantetheine attachment site | 1211 | 1226 | - |
| Gene3D | G3DSA:2.40.110.10 | 739 | 844 | 2.1E-83 | |||
| SUPERFAMILY | SSF56801 | 7 | 575 | 3.27E-107 | |||
| CDD | cd00567 | ACAD | 711 | 991 | 2.59276E-64 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.